HuR/methyl-HuR and AUF1 regulate the MAT expressed during liver proliferation, differentiation, and carcinogenesis
- PMID: 20102719
- PMCID: PMC2860011
- DOI: 10.1053/j.gastro.2010.01.032
HuR/methyl-HuR and AUF1 regulate the MAT expressed during liver proliferation, differentiation, and carcinogenesis
Abstract
Background & aims: Hepatic de-differentiation, liver development, and malignant transformation are processes in which the levels of hepatic S-adenosylmethionine are tightly regulated by 2 genes: methionine adenosyltransferase 1A (MAT1A) and methionine adenosyltransferase 2A (MAT2A). MAT1A is expressed in the adult liver, whereas MAT2A expression primarily is extrahepatic and is associated strongly with liver proliferation. The mechanisms that regulate these expression patterns are not completely understood.
Methods: In silico analysis of the 3' untranslated region of MAT1A and MAT2A revealed putative binding sites for the RNA-binding proteins AU-rich RNA binding factor 1 (AUF1) and HuR, respectively. We investigated the posttranscriptional regulation of MAT1A and MAT2A by AUF1, HuR, and methyl-HuR in the aforementioned biological processes.
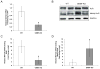
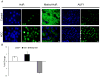
Results: During hepatic de-differentiation, the switch between MAT1A and MAT2A coincided with an increase in HuR and AUF1 expression. S-adenosylmethionine treatment altered this homeostasis by shifting the balance of AUF1 and methyl-HuR/HuR, which was identified as an inhibitor of MAT2A messenger RNA (mRNA) stability. We also observed a similar temporal distribution and a functional link between HuR, methyl-HuR, AUF1, and MAT1A and MAT2A during fetal liver development. Immunofluorescent analysis revealed increased levels of HuR and AUF1, and a decrease in methyl-HuR levels in human livers with hepatocellular carcinoma (HCC).
Conclusions: Our data strongly support a role for AUF1 and HuR/methyl-HuR in liver de-differentiation, development, and human HCC progression through the posttranslational regulation of MAT1A and MAT2A mRNAs.
Copyright 2010 AGA Institute. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
None of the authors have conflict of interest to declare
Figures





Similar articles
-
Pleiotropic effects of methionine adenosyltransferases deregulation as determinants of liver cancer progression and prognosis.J Hepatol. 2013 Oct;59(4):830-41. doi: 10.1016/j.jhep.2013.04.031. Epub 2013 May 7. J Hepatol. 2013. PMID: 23665184 Review.
-
Role of transcriptional and posttranscriptional regulation of methionine adenosyltransferases in liver cancer progression.Hepatology. 2012 Jul;56(1):165-75. doi: 10.1002/hep.25643. Epub 2012 Jun 5. Hepatology. 2012. PMID: 22318685
-
Polyamines regulate the stability of JunD mRNA by modulating the competitive binding of its 3' untranslated region to HuR and AUF1.Mol Cell Biol. 2010 Nov;30(21):5021-32. doi: 10.1128/MCB.00807-10. Epub 2010 Aug 30. Mol Cell Biol. 2010. PMID: 20805360 Free PMC article.
-
Interaction between 3' untranslated region of calcitonin receptor messenger ribonucleic acid (RNA) and adenylate/uridylate (AU)-rich element binding proteins (AU-rich RNA-binding factor 1 and Hu antigen R).Endocrinology. 2004 Apr;145(4):1730-8. doi: 10.1210/en.2003-0862. Epub 2004 Jan 8. Endocrinology. 2004. PMID: 14715706
-
Methionine adenosyltransferases in liver cancer.World J Gastroenterol. 2019 Aug 21;25(31):4300-4319. doi: 10.3748/wjg.v25.i31.4300. World J Gastroenterol. 2019. PMID: 31496615 Free PMC article. Review.
Cited by
-
Hepatic levels of S-adenosylmethionine regulate the adaptive response to fasting.Cell Metab. 2023 Aug 8;35(8):1373-1389.e8. doi: 10.1016/j.cmet.2023.07.002. Epub 2023 Jul 31. Cell Metab. 2023. PMID: 37527658 Free PMC article.
-
RNA-binding proteins and their role in kidney disease.Nat Rev Nephrol. 2022 Mar;18(3):153-170. doi: 10.1038/s41581-021-00497-1. Epub 2021 Nov 3. Nat Rev Nephrol. 2022. PMID: 34732838 Review.
-
MicroRNA-203 impacts on the growth, aggressiveness and prognosis of hepatocellular carcinoma by targeting MAT2A and MAT2B genes.Oncotarget. 2019 Apr 19;10(29):2835-2854. doi: 10.18632/oncotarget.26838. eCollection 2019 Apr 19. Oncotarget. 2019. PMID: 31073374 Free PMC article.
-
Role of 3'-untranslated region translational control in cancer development, diagnostics and treatment.World J Biol Chem. 2014 Feb 26;5(1):40-57. doi: 10.4331/wjbc.v5.i1.40. World J Biol Chem. 2014. PMID: 24600513 Free PMC article. Review.
-
AU-binding factor 1 expression was correlated with metadherin expression and progression of hepatocellular carcinoma.Tumour Biol. 2014 Mar;35(3):2747-51. doi: 10.1007/s13277-013-1362-2. Epub 2013 Nov 9. Tumour Biol. 2014. PMID: 24213928
References
-
- Mato JM, Corrales FJ, Lu SC, et al. S-Adenosylmethionine: a control switch that regulates liver function. FASEB J. 2002;16:15–26. - PubMed
-
- Gil B, Casado M, Pajares M, et al. Differential expression pattern of methionine adenosyltransferase isoenzymes during rat liver development. Hepatology. 1996;24:876–881. - PubMed
-
- Cai J, Mao Z, Hwang JJ, et al. Differential expression of methionine adenosyltransferase genes influences the rate of growth of human hepatocellular carcinoma cells. Cancer Res. 1998;58:1444–1450. - PubMed
-
- Pascale RM, Marras V, Simile MM, et al. Chemoprevention of rat liver carcinogenesis by S-adenosyl-L-methionine: A long-term study. Cancer Res. 1992;52:4979–4986. - PubMed
-
- Shivapurkar N, Poirier LA. Tissue levels of S-adenosylmethionine and S- adenosylhomocysteine in rats fed methyl-deficiency, amino acid-defined diets for one to five weeks. Carcinogenesis. 1983;4:1051–1057. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

