Comparison of the biological and biochemical activities of several members of the alphaherpesvirus ICP0 family of proteins
- PMID: 20106921
- PMCID: PMC2838103
- DOI: 10.1128/JVI.02544-09
Comparison of the biological and biochemical activities of several members of the alphaherpesvirus ICP0 family of proteins
Abstract
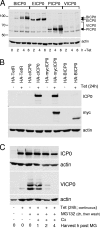
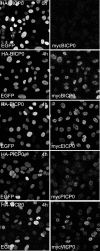
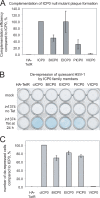
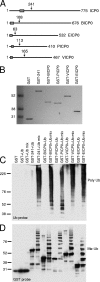
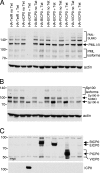
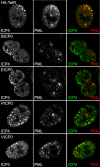
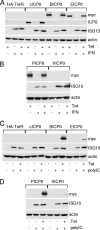
Immediate-early protein ICP0 of herpes simplex virus type 1 (HSV-1) is an E3 ubiquitin ligase of the RING finger class that is required for efficient lytic infection and reactivation from latency. Other alphaherpesviruses also express ICP0-related RING finger proteins, but these have limited homology outside the core RING domain. Existing evidence indicates that ICP0 family members have similar properties, but there has been no systematic comparison of the biochemical activities and biological functions of these proteins. Here, we describe an inducible cell line system that allows expression of the ICP0-related proteins of bovine herpes virus type 1 (BHV-1), equine herpesvirus type 1 (EHV-1), pseudorabies virus (PRV), and varicella-zoster virus (VZV) and their subsequent functional analysis. We report that the RING domains of all the proteins have E3 ubiquitin ligase activity in vitro. The BHV-1, EHV-1, and PRV proteins complement ICP0-null mutant HSV-1 plaque formation and induce derepression of quiescent HSV-1 genomes to levels similar to those achieved by ICP0 itself. VICP0, the ICP0 expressed by VZV, was found to be extremely unstable, which limited its analysis in this system. We compared the abilities of the ICP0-related proteins to disrupt ND10, to induce degradation of PML and Sp100, to affect key components of the interferon signaling pathway, and to interfere with induction of interferon-stimulated genes. We found that the property that correlated most closely with their biological activities was the ability to preclude the recruitment of cellular ND10 proteins to sites closely associated with incoming HSV-1 genomes and early replication compartments.
Figures



Similar articles
-
The HSV-1 ubiquitin ligase ICP0: Modifying the cellular proteome to promote infection.Virus Res. 2020 Aug;285:198015. doi: 10.1016/j.virusres.2020.198015. Epub 2020 May 13. Virus Res. 2020. PMID: 32416261 Free PMC article. Review.
-
Two overlapping regions within the N-terminal half of the herpes simplex virus 1 E3 ubiquitin ligase ICP0 facilitate the degradation and dissociation of PML and dissociation of Sp100 from ND10.J Virol. 2013 Dec;87(24):13287-96. doi: 10.1128/JVI.02304-13. Epub 2013 Oct 2. J Virol. 2013. PMID: 24089549 Free PMC article.
-
Alphaherpesvirus proteins related to herpes simplex virus type 1 ICP0 affect cellular structures and proteins.J Virol. 2000 Nov;74(21):10006-17. doi: 10.1128/jvi.74.21.10006-10017.2000. J Virol. 2000. PMID: 11024129 Free PMC article.
-
Replication of ICP0-null mutant herpes simplex virus type 1 is restricted by both PML and Sp100.J Virol. 2008 Mar;82(6):2661-72. doi: 10.1128/JVI.02308-07. Epub 2007 Dec 26. J Virol. 2008. PMID: 18160441 Free PMC article.
-
Regulation of alphaherpesvirus infections by the ICP0 family of proteins.J Gen Virol. 2013 Mar;94(Pt 3):465-481. doi: 10.1099/vir.0.048900-0. Epub 2012 Dec 12. J Gen Virol. 2013. PMID: 23239572 Review.
Cited by
-
Crystal Structure of USP7 Ubiquitin-like Domains with an ICP0 Peptide Reveals a Novel Mechanism Used by Viral and Cellular Proteins to Target USP7.PLoS Pathog. 2015 Jun 5;11(6):e1004950. doi: 10.1371/journal.ppat.1004950. eCollection 2015 Jun. PLoS Pathog. 2015. PMID: 26046769 Free PMC article.
-
Sp140L functions as a herpesvirus restriction factor suppressing viral transcription and activating interferon-stimulated genes.Proc Natl Acad Sci U S A. 2025 Jun 24;122(25):e2426339122. doi: 10.1073/pnas.2426339122. Epub 2025 Jun 17. Proc Natl Acad Sci U S A. 2025. PMID: 40526717
-
High-yield expression in Escherichia coli and purification of mouse ubiquitin-activating enzyme E1.Mol Biotechnol. 2012 Jul;51(3):254-61. doi: 10.1007/s12033-011-9463-x. Mol Biotechnol. 2012. PMID: 22012022
-
Evaluation of the effects of acyclovir and/or human amniotic membrane on herpes virus culture and quantitative virus inactivity by real-time polymerase chain reaction.Int J Ophthalmol. 2014 Aug 18;7(4):626-31. doi: 10.3980/j.issn.2222-3959.2014.04.07. eCollection 2014. Int J Ophthalmol. 2014. PMID: 25161932 Free PMC article.
-
The HSV-1 ubiquitin ligase ICP0: Modifying the cellular proteome to promote infection.Virus Res. 2020 Aug;285:198015. doi: 10.1016/j.virusres.2020.198015. Epub 2020 May 13. Virus Res. 2020. PMID: 32416261 Free PMC article. Review.
References
-
- Barlow, P. N., B. Luisi, A. Milner, M. Elliott, and R. Everett. 1994. Structure of the C3HC4 domain by 1H-nuclear magnetic resonance spectroscopy. A new structural class of zinc-finger. J. Mol. Biol. 237:201-211. - PubMed
-
- Boldogkoi, Z., A. Braun, and I. Fodor. 2000. Replication and virulence of early protein 0 and long latency transcript deficient mutants of the Aujeszky's disease (pseudorabies) virus. Microbes Infect. 2:1321-1328. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

