The submergence tolerance regulator Sub1A mediates stress-responsive expression of AP2/ERF transcription factors
- PMID: 20107022
- PMCID: PMC2832257
- DOI: 10.1104/pp.109.152157
The submergence tolerance regulator Sub1A mediates stress-responsive expression of AP2/ERF transcription factors
Abstract
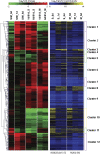
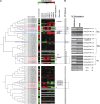
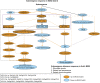
We previously characterized the rice (Oryza sativa) Submergence1 (Sub1) locus encoding three ethylene-responsive factor (ERF) transcriptional regulators. Genotypes carrying the Sub1A-1 allele are tolerant of prolonged submergence. To elucidate the mechanism of Sub1A-1-mediated tolerance, we performed transcriptome analyses comparing the temporal submergence response of Sub1A-1-containing tolerant M202(Sub1) with the intolerant isoline M202 lacking this gene. We identified 898 genes displaying Sub1A-1-dependent regulation. Integration of the expression data with publicly available metabolic pathway data identified submergence tolerance-associated pathways governing anaerobic respiration, hormone responses, and antioxidant systems. Of particular interest were a set of APETALA2 (AP2)/ERF family transcriptional regulators that are associated with the Sub1A-1-mediated response upon submergence. Visualization of expression patterns of the AP2/ERF superfamily members in a phylogenetic context resolved 12 submergence-regulated AP2/ERFs into three putative functional groups: (1) anaerobic respiration and cytokinin-mediated delay in senescence via ethylene accumulation during submergence (three ERFs); (2) negative regulation of ethylene-dependent gene expression (five ERFs); and (3) negative regulation of gibberellin-mediated shoot elongation (four ERFs). These results confirm that the presence of Sub1A-1 impacts multiple pathways of response to submergence.
Figures




Similar articles
-
A variable cluster of ethylene response factor-like genes regulates metabolic and developmental acclimation responses to submergence in rice.Plant Cell. 2006 Aug;18(8):2021-34. doi: 10.1105/tpc.106.043000. Epub 2006 Jun 30. Plant Cell. 2006. PMID: 16816135 Free PMC article.
-
The submergence tolerance regulator SUB1A mediates crosstalk between submergence and drought tolerance in rice.Plant Cell. 2011 Jan;23(1):412-27. doi: 10.1105/tpc.110.080325. Epub 2011 Jan 14. Plant Cell. 2011. PMID: 21239643 Free PMC article.
-
SUB1A-mediated submergence tolerance response in rice involves differential regulation of the brassinosteroid pathway.New Phytol. 2013 Jun;198(4):1060-1070. doi: 10.1111/nph.12202. Epub 2013 Mar 18. New Phytol. 2013. PMID: 23496140
-
Unraveling the genetic enigma of rice submergence tolerance: Shedding light on the role of ethylene response factor-encoding gene SUB1A-1.Plant Physiol Biochem. 2024 Jan;206:108224. doi: 10.1016/j.plaphy.2023.108224. Epub 2023 Nov 25. Plant Physiol Biochem. 2024. PMID: 38091930 Review.
-
[Submergence tolerance and Sub1 locus in rice].Yi Chuan. 2010 Sep;32(9):886-92. Yi Chuan. 2010. PMID: 20870609 Review. Chinese.
Cited by
-
Improvement of enzymatic saccharification yield in Arabidopsis thaliana by ectopic expression of the rice SUB1A-1 transcription factor.PeerJ. 2015 Mar 3;3:e817. doi: 10.7717/peerj.817. eCollection 2015. PeerJ. 2015. PMID: 25780769 Free PMC article.
-
A Positive Feedback Loop Governed by SUB1A1 Interaction with MITOGEN-ACTIVATED PROTEIN KINASE3 Imparts Submergence Tolerance in Rice.Plant Cell. 2016 May;28(5):1127-43. doi: 10.1105/tpc.15.01001. Epub 2016 Apr 14. Plant Cell. 2016. PMID: 27081183 Free PMC article.
-
The Use of DNA Markers in Rice Breeding for Blast Resistance and Submergence Tolerance as a Weed Control Factor.Plants (Basel). 2025 Jun 13;14(12):1815. doi: 10.3390/plants14121815. Plants (Basel). 2025. PMID: 40573803 Free PMC article.
-
Understanding AP2/ERF Transcription Factor Responses and Tolerance to Various Abiotic Stresses in Plants: A Comprehensive Review.Int J Mol Sci. 2024 Jan 11;25(2):893. doi: 10.3390/ijms25020893. Int J Mol Sci. 2024. PMID: 38255967 Free PMC article. Review.
-
Genetic and molecular characterization of submergence response identifies Subtol6 as a major submergence tolerance locus in maize.PLoS One. 2015 Mar 25;10(3):e0120385. doi: 10.1371/journal.pone.0120385. eCollection 2015. PLoS One. 2015. PMID: 25806518 Free PMC article.
References
-
- Almeida AM, Vriezen WH, Van der Straeten D. (2003) Molecular and physiological mechanisms of flooding avoidance and tolerance in rice. Russ J Plant Physiol 50: 743–751
-
- Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, et al. (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301: 653–657 - PubMed
-
- Bailey-Serres J, Voesenek LA. (2008) Flooding stress: acclimations and genetic diversity. Annu Rev Plant Biol 59: 313–339 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

