Widespread and tissue specific age-related DNA methylation changes in mice
- PMID: 20107151
- PMCID: PMC2840983
- DOI: 10.1101/gr.096826.109
Widespread and tissue specific age-related DNA methylation changes in mice
Abstract
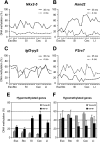
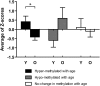
Aberrant methylation of promoter CpG islands in cancer is associated with silencing of tumor-suppressor genes, and age-dependent hypermethylation in normal appearing mucosa may be a risk factor for human colon cancer. It is not known whether this age-related DNA methylation phenomenon is specific to human tissues. We performed comprehensive DNA methylation profiling of promoter regions in aging mouse intestine using methylated CpG island amplification in combination with microarray analysis. By comparing C57BL/6 mice at 3-mo-old versus 35-mo-old for 3627 detectable autosomal genes, we found 774 (21%) that showed increased methylation and 466 (13%) that showed decreased methylation. We used pyrosequencing to quantitatively validate the microarray data and confirmed linear age-related methylation changes for all 12 genomic regions examined. We then examined 11 changed genomic loci for age-related methylation in other tissues. Of these, three of 11 showed similar changes in lung, seven of 11 changed in liver, and six of 11 changed in spleen, though to a lower degree than the changes seen in colon. There was partial conservation between age-related hypermethylation in human and mouse intestines, and Polycomb targets in embryonic stem cells were enriched among the hypermethylated genes. Our findings demonstrate a surprisingly high rate of hyper- and hypomethylation as a function of age in normal mouse small intestine tissues and a strong tissue-specificity to the process. We conclude that epigenetic deregulation is a common feature of aging in mammals.
Figures


References
-
- Ahuja N, Issa JP. Aging, methylation and cancer. Histol Histopathol. 2000;15:835–842. - PubMed
-
- Ahuja N, Li Q, Mohan AL, Baylin SB, Issa JP. Aging and DNA methylation in colorectal mucosa and cancer. Cancer Res. 1998;58:5489–5494. - PubMed
-
- Bird AP. CpG-rich islands and the function of DNA methylation. Nature. 1986;321:209–213. - PubMed
-
- Boyer LA, Plath K, Zeitlinger J, Brambrink T, Medeiros LA, Lee TI, Levine SS, Wernig M, Tajonar A, Ray MK, et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature. 2006;441:349–353. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
