Persistent ER stress induces the spliced leader RNA silencing pathway (SLS), leading to programmed cell death in Trypanosoma brucei
- PMID: 20107599
- PMCID: PMC2809764
- DOI: 10.1371/journal.ppat.1000731
Persistent ER stress induces the spliced leader RNA silencing pathway (SLS), leading to programmed cell death in Trypanosoma brucei
Abstract
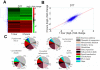
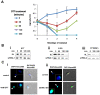
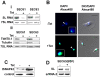
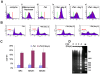
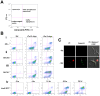
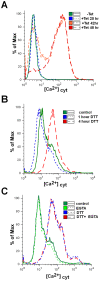
Trypanosomes are parasites that cycle between the insect host (procyclic form) and mammalian host (bloodstream form). These parasites lack conventional transcription regulation, including factors that induce the unfolded protein response (UPR). However, they possess a stress response mechanism, the spliced leader RNA silencing (SLS) pathway. SLS elicits shut-off of spliced leader RNA (SL RNA) transcription by perturbing the binding of the transcription factor tSNAP42 to its cognate promoter, thus eliminating trans-splicing of all mRNAs. Induction of endoplasmic reticulum (ER) stress in procyclic trypanosomes elicits changes in the transcriptome similar to those induced by conventional UPR found in other eukaryotes. The mechanism of up-regulation under ER stress is dependent on differential stabilization of mRNAs. The transcriptome changes are accompanied by ER dilation and elevation in the ER chaperone, BiP. Prolonged ER stress induces SLS pathway. RNAi silencing of SEC63, a factor that participates in protein translocation across the ER membrane, or SEC61, the translocation channel, also induces SLS. Silencing of these genes or prolonged ER stress led to programmed cell death (PCD), evident by exposure of phosphatidyl serine, DNA laddering, increase in reactive oxygen species (ROS) production, increase in cytoplasmic Ca(2+), and decrease in mitochondrial membrane potential, as well as typical morphological changes observed by transmission electron microscopy (TEM). ER stress response is also induced in the bloodstream form and if the stress persists it leads to SLS. We propose that prolonged ER stress induces SLS, which serves as a unique death pathway, replacing the conventional caspase-mediated PCD observed in higher eukaryotes.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


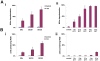

Similar articles
-
Induction of ER stress response leading to programmed cell death in Trypanosoma brucei.Methods Enzymol. 2011;489:189-205. doi: 10.1016/B978-0-12-385116-1.00011-X. Methods Enzymol. 2011. PMID: 21266231
-
Spliced leader RNA silencing (SLS) - a programmed cell death pathway in Trypanosoma brucei that is induced upon ER stress.Parasit Vectors. 2012 May 31;5:107. doi: 10.1186/1756-3305-5-107. Parasit Vectors. 2012. PMID: 22650251 Free PMC article. Review.
-
The response of trypanosomes and other eukaryotes to ER stress and the spliced leader RNA silencing (SLS) pathway in Trypanosoma brucei.Crit Rev Biochem Mol Biol. 2015;50(3):256-67. doi: 10.3109/10409238.2015.1042541. Epub 2015 May 19. Crit Rev Biochem Mol Biol. 2015. PMID: 25985970 Review.
-
Transcriptome and proteome analyses and the role of atypical calpain protein and autophagy in the spliced leader silencing pathway in Trypanosoma brucei.Mol Microbiol. 2016 Oct;102(1):1-21. doi: 10.1111/mmi.13417. Epub 2016 Aug 31. Mol Microbiol. 2016. PMID: 27161313
-
The Spliced Leader RNA Silencing (SLS) Pathway in Trypanosoma brucei Is Induced by Perturbations of Endoplasmic Reticulum, Golgi Complex, or Mitochondrial Protein Factors: Functional Analysis of SLS-Inducing Kinase PK3.mBio. 2021 Dec 21;12(6):e0260221. doi: 10.1128/mBio.02602-21. Epub 2021 Nov 30. mBio. 2021. PMID: 34844425 Free PMC article.
Cited by
-
The endoplasmic reticulum of trypanosomatids: An unrevealed road for chemotherapy.Front Cell Infect Microbiol. 2022 Nov 10;12:1057774. doi: 10.3389/fcimb.2022.1057774. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 36439218 Free PMC article. Review.
-
Autophagy in Trypanosoma brucei: amino acid requirement and regulation during different growth phases.PLoS One. 2014 Apr 3;9(4):e93875. doi: 10.1371/journal.pone.0093875. eCollection 2014. PLoS One. 2014. PMID: 24699810 Free PMC article.
-
Benzoxaborole treatment perturbs S-adenosyl-L-methionine metabolism in Trypanosoma brucei.PLoS Negl Trop Dis. 2018 May 14;12(5):e0006450. doi: 10.1371/journal.pntd.0006450. eCollection 2018 May. PLoS Negl Trop Dis. 2018. PMID: 29758036 Free PMC article.
-
Blocking variant surface glycoprotein synthesis alters endoplasmic reticulum exit sites/Golgi homeostasis in Trypanosoma brucei.Traffic. 2018 Jun;19(6):391-405. doi: 10.1111/tra.12561. Epub 2018 Apr 6. Traffic. 2018. PMID: 29533496 Free PMC article.
-
Intracellular eukaryotic parasites have a distinct unfolded protein response.PLoS One. 2011 Apr 29;6(4):e19118. doi: 10.1371/journal.pone.0019118. PLoS One. 2011. PMID: 21559456 Free PMC article.
References
-
- Travers KJ, Patil CK, Wodicka L, Lockhart DJ, Weissman JS, et al. Functional and genomic analyses reveal an essential coordination between the unfolded protein response and ER-associated degradation. Cell. 2000;101:249–258. - PubMed
-
- Ron D, Walter P. Signal integration in the endoplasmic reticulum unfolded protein response. Nat Rev Mol Cell Biol. 2007;8:519–529. - PubMed
-
- Bernales S, Papa FR, Walter P. Intracellular signaling by the unfolded protein response. Annu Rev Cell Dev Biol. 2006;22:487–508. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

