ATRX interacts with H3.3 in maintaining telomere structural integrity in pluripotent embryonic stem cells
- PMID: 20110566
- PMCID: PMC2840985
- DOI: 10.1101/gr.101477.109
ATRX interacts with H3.3 in maintaining telomere structural integrity in pluripotent embryonic stem cells
Abstract
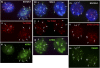
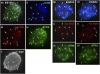
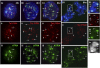
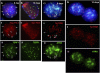
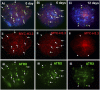
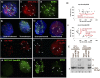
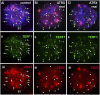
ATRX (alpha thalassemia/mental retardation syndrome X-linked) belongs to the SWI2/SNF2 family of chromatin remodeling proteins. Besides the ATPase/helicase domain at its C terminus, it contains a PHD-like zinc finger at the N terminus. Mutations in the ATRX gene are associated with X-linked mental retardation (XLMR) often accompanied by alpha thalassemia (ATRX syndrome). Although ATRX has been postulated to be a transcriptional regulator, its precise roles remain undefined. We demonstrate ATRX localization at the telomeres in interphase mouse embryonic stem (ES) cells in synchrony with the incorporation of H3.3 during telomere replication at S phase. Moreover, we found that chromobox homolog 5 (CBX5) (also known as heterochromatin protein 1 alpha, or HP1 alpha) is also present at the telomeres in ES cells. We show by coimmunoprecipitation that this localization is dependent on the association of ATRX with histone H3.3, and that mutating the K4 residue of H3.3 significantly diminishes ATRX and H3.3 interaction. RNAi-knockdown of ATRX induces a telomere-dysfunction phenotype and significantly reduces CBX5 enrichment at the telomeres. These findings suggest a novel function of ATRX, working in conjunction with H3.3 and CBX5, as a key regulator of ES-cell telomere chromatin.
Figures


References
-
- Berube NG, Jagla M, Smeenk C, De Repentigny Y, Kothary R, Picketts DJ. Neurodevelopmental defects resulting from ATRX overexpression in transgenic mice. Hum Mol Genet. 2002;11:253–261. - PubMed
-
- Bienz M. The PHD finger, a nuclear protein-interaction domain. Trends Biochem Sci. 2006;31:35–40. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
