Changes in gene expression in regions of the extended amygdala of alcohol-preferring rats after binge-like alcohol drinking
- PMID: 20116196
- PMCID: PMC2831121
- DOI: 10.1016/j.alcohol.2009.12.001
Changes in gene expression in regions of the extended amygdala of alcohol-preferring rats after binge-like alcohol drinking
Abstract
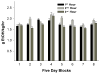
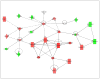
The objective of this study was to determine time-course changes in gene expression within two regions of the extended amygdala after binge-like alcohol drinking by alcohol-preferring (P) rats. Adult male P rats were given 1-h access to 15 and 30% ethanol three times daily for 8 weeks. Rats (n = 10/time point for ethanol and n = 6/time point for water) were killed by decapitation 1, 6, and 24 h after the last drinking episode. RNA was prepared from individual micropunch samples of the nucleus accumbens shell (ACB-shell) and central nucleus of the amygdala (CeA); analyses were conducted with Affymetrix Rat Genome 230.2 GeneChips. Ethanol intakes were 1.5-2 g/kg for each of the three sessions. There were no genes that were statistically different between the ethanol and water control groups at any individual time point. Therefore, an overall effect, comparing the water control and ethanol groups, was determined. In the ACB-shell and CeA, there were 276 and 402 probe sets for named genes, respectively, that differed between the two groups. There were 1.5-3.6-fold more genes with increased expression than with decreased expression in the ethanol-drinking group, with most differences between 1.1- and 1.2-fold. Among the differences between the ethanol and water control groups were several significant biological processes categories that were in common between the two regions (e.g., synaptic transmission, neurite development); however, within these categories, there were few genes in common between the two regions. Overall, the results indicate that binge-like alcohol drinking by P rats produces region-dependent changes in the expression of genes that could alter transcription, synaptic function, and neuronal plasticity in the ACB-shell and CeA; within each region, different mechanisms may underlie these alterations because there were few common ethanol-responsive genes between the ACB-shell and CeA.
2010 Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Changes in gene expression within the extended amygdala following binge-like alcohol drinking by adolescent alcohol-preferring (P) rats.Pharmacol Biochem Behav. 2014 Feb;117:52-60. doi: 10.1016/j.pbb.2013.12.009. Epub 2013 Dec 16. Pharmacol Biochem Behav. 2014. PMID: 24355552 Free PMC article.
-
Gene expression within the extended amygdala of 5 pairs of rat lines selectively bred for high or low ethanol consumption.Alcohol. 2013 Nov;47(7):517-29. doi: 10.1016/j.alcohol.2013.08.004. Epub 2013 Oct 1. Alcohol. 2013. PMID: 24157127 Free PMC article.
-
Differential gene expression in the nucleus accumbens with ethanol self-administration in inbred alcohol-preferring rats.Pharmacol Biochem Behav. 2008 Jun;89(4):481-98. doi: 10.1016/j.pbb.2008.01.023. Epub 2008 Feb 26. Pharmacol Biochem Behav. 2008. PMID: 18405950 Free PMC article.
-
Central nucleus of the amygdala and the effects of alcohol and alcohol-drinking behavior in rodents.Pharmacol Biochem Behav. 2002 Mar;71(3):509-15. doi: 10.1016/s0091-3057(01)00680-3. Pharmacol Biochem Behav. 2002. PMID: 11830185 Review.
-
[Epigenetic mechanisms and alcohol use disorders: a potential therapeutic target].Biol Aujourdhui. 2017;211(1):83-91. doi: 10.1051/jbio/2017014. Epub 2017 Jul 6. Biol Aujourdhui. 2017. PMID: 28682229 Review. French.
Cited by
-
Gene expression in brain and liver produced by three different regimens of alcohol consumption in mice: comparison with immune activation.PLoS One. 2013;8(3):e59870. doi: 10.1371/journal.pone.0059870. Epub 2013 Mar 29. PLoS One. 2013. PMID: 23555817 Free PMC article.
-
The Role of Orexin Signaling in the Ventral Tegmental Area and Central Amygdala in Modulating Binge-Like Ethanol Drinking Behavior.Alcohol Clin Exp Res. 2017 Mar;41(3):551-561. doi: 10.1111/acer.13336. Epub 2017 Feb 9. Alcohol Clin Exp Res. 2017. PMID: 28097729 Free PMC article.
-
Rodent models and mechanisms of voluntary binge-like ethanol consumption: Examples, opportunities, and strategies for preclinical research.Prog Neuropsychopharmacol Biol Psychiatry. 2016 Feb 4;65:297-308. doi: 10.1016/j.pnpbp.2015.05.012. Epub 2015 Jun 2. Prog Neuropsychopharmacol Biol Psychiatry. 2016. PMID: 26021391 Free PMC article. Review.
-
Dynamic regulation of CeA gene expression during acute and protracted abstinence from chronic binge drinking of male and female C57BL/6J mice.bioRxiv [Preprint]. 2024 Feb 4:2024.02.02.578650. doi: 10.1101/2024.02.02.578650. bioRxiv. 2024. Update in: Alcohol. 2024 Nov;120:179-193. doi: 10.1016/j.alcohol.2024.06.005. PMID: 38352404 Free PMC article. Updated. Preprint.
-
Binge Ethanol Drinking Produces Sexually Divergent and Distinct Changes in Nucleus Accumbens Signaling Cascades and Pathways in Adult C57BL/6J Mice.Front Genet. 2018 Sep 10;9:325. doi: 10.3389/fgene.2018.00325. eCollection 2018. Front Genet. 2018. PMID: 30250478 Free PMC article.
References
-
- Al-Housseini AM, Sivanandam TM, Bradbury EL, Tannenberg RK, Dodd PR, Gu Q. Upregulation of β-catenin levels in superior frontal cortex of chronic alcoholics. Alcohol Clin Exp Res. 2008;32:1080–1090. - PubMed
-
- Alexander-Kaufman K, James G, Sheedy D, Harper C, Matsumoto I. Differential protein expression in the prefrontal white matter of human alcoholics: a proteomics study. Molecular Psych. 2006;11:56–65. - PubMed
-
- Alexander-Kaufman K, Harper C, Wilce P, Matsumoto I. Cerebellar vermis proteome of chronic alcoholic individuals. Alcohol Clin Exp Res. 2007;31:1286–1296. - PubMed
-
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–29. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- U01 AA013521/AA/NIAAA NIH HHS/United States
- R24 AA015512/AA/NIAAA NIH HHS/United States
- U01 AA016660/AA/NIAAA NIH HHS/United States
- AA013521/AA/NIAAA NIH HHS/United States
- U01 AA016652/AA/NIAAA NIH HHS/United States
- P60 AA007611/AA/NIAAA NIH HHS/United States
- P50 AA007611/AA/NIAAA NIH HHS/United States
- AA07611/AA/NIAAA NIH HHS/United States
- U01 AA013522/AA/NIAAA NIH HHS/United States
- AA013522/AA/NIAAA NIH HHS/United States
- U24 AA013522/AA/NIAAA NIH HHS/United States
- AA016660/AA/NIAAA NIH HHS/United States
- AA016652/AA/NIAAA NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

