Histidine kinase-mediated production and autoassembly of Porphyromonas gingivalis fimbriae
- PMID: 20118268
- PMCID: PMC2838039
- DOI: 10.1128/JB.01474-09
Histidine kinase-mediated production and autoassembly of Porphyromonas gingivalis fimbriae
Abstract
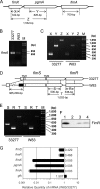
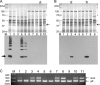
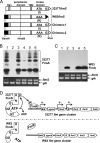
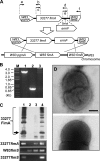
Porphyromonas gingivalis, a Gram-negative oral anaerobe, is strongly associated with chronic adult periodontitis, and it utilizes FimA fimbriae to persistently colonize and evade host defenses in the periodontal crevice. The FimA-related gene cluster (the fim gene cluster) is positively regulated by the FimS-FimR two-component system. In this study, comparative analyses between fimbriate type strain ATCC 33277 and fimbria-deficient strain W83 revealed differences in their fimS loci, which encode FimS histidine kinase. Using a reciprocal gene exchange system, we established that FimS from W83 is malfunctional. Complementation analysis with chimeric fimS constructs revealed that W83 FimS has a defective kinase domain due to a truncated conserved G3 box motif that provides an ATP-binding pocket. The introduction of the functional fimS from 33277 restored the production, but not polymerization, of endogenous FimA subunits in W83. Further analyses with a fimA-exchanged W83 isogenic strain showed that even the fimbria-deficient W83 retains the ability to polymerize FimA from 33277, indicating the assembly of mature FimA by a primary structure-dependent mechanism. It also was shown that the substantial expression of 33277-type FimA fimbriae in the W83 derivative requires the introduction and expression of the functional 33277 fimS. These findings indicate that FimSR is the unique and universal regulatory system that activates the fim gene cluster in a fimA genotype-independent manner.
Figures



Similar articles
-
A regulation cascade controls expression of Porphyromonas gingivalis fimbriae via the FimR response regulator.Mol Microbiol. 2004 Oct;54(2):546-60. doi: 10.1111/j.1365-2958.2004.04291.x. Mol Microbiol. 2004. PMID: 15469523
-
Identification of Critical Amino Acid Residues of a Two-Component Sensor Protein for Signal Sensing in Porphyromonas gingivalis Fimbriation via Random Mutant Library Construction.Pathogens. 2024 Apr 10;13(4):309. doi: 10.3390/pathogens13040309. Pathogens. 2024. PMID: 38668264 Free PMC article.
-
Porphyromonas gingivalis short fimbriae are regulated by a FimS/FimR two-component system.FEMS Microbiol Lett. 2007 Jun;271(2):214-21. doi: 10.1111/j.1574-6968.2007.00722.x. Epub 2007 Apr 20. FEMS Microbiol Lett. 2007. PMID: 17451448 Free PMC article.
-
[FimA fimbriae of the periodontal disease-associated bacterium Porphyromonas gingivalis].Yakugaku Zasshi. 2013;133(9):963-74. doi: 10.1248/yakushi.13-00177. Yakugaku Zasshi. 2013. PMID: 23995804 Review. Japanese.
-
Molecular interaction of Porphyromonas gingivalis with host cells: implication for the microbial pathogenesis of periodontal disease.J Periodontol. 2003 Jan;74(1):90-6. doi: 10.1902/jop.2003.74.1.90. J Periodontol. 2003. PMID: 12593602 Review.
Cited by
-
A two-component system regulates gene expression of the type IX secretion component proteins via an ECF sigma factor.Sci Rep. 2016 Mar 21;6:23288. doi: 10.1038/srep23288. Sci Rep. 2016. PMID: 26996145 Free PMC article.
-
VimA-dependent modulation of acetyl coenzyme A levels and lipid A biosynthesis can alter virulence in Porphyromonas gingivalis.Infect Immun. 2012 Feb;80(2):550-64. doi: 10.1128/IAI.06062-11. Epub 2011 Dec 5. Infect Immun. 2012. PMID: 22144476 Free PMC article.
-
Phylogenetic diversity in fim and mfa gene clusters between Porphyromonas gingivalis and Porphyromonas gulae, as a potential cause of host specificity.J Oral Microbiol. 2020 Jun 19;12(1):1775333. doi: 10.1080/20002297.2020.1775333. J Oral Microbiol. 2020. PMID: 32944148 Free PMC article.
-
Roles of Porphyromonas gingivalis and its virulence factors in periodontitis.Adv Protein Chem Struct Biol. 2020;120:45-84. doi: 10.1016/bs.apcsb.2019.12.001. Epub 2020 Jan 10. Adv Protein Chem Struct Biol. 2020. PMID: 32085888 Free PMC article. Review.
-
Deletion of lipoprotein PG0717 in Porphyromonas gingivalis W83 reduces gingipain activity and alters trafficking in and response by host cells.PLoS One. 2013 Sep 12;8(9):e74230. doi: 10.1371/journal.pone.0074230. eCollection 2013. PLoS One. 2013. PMID: 24069284 Free PMC article.
References
-
- Akiyama, Y., K. Onizuka, T. Noguchi, and M. Ando. 1998. Parallel protein information analysis (PAPIA) system running on a 64-node PC cluster, p. 131-140. In Proc. 9th Genome Informatics Workshop (GIW'98), Universal Academy Press, Tokyo, Japan. - PubMed
-
- Belton, C. M., K. T. Izutsu, P. C. Goodwin, Y. Park, and R. J. Lamont. 1999. Fluorescence image analysis of the association between Porphyromonas gingivalis and gingival epithelial cells. Cell. Microbiol. 1:215-223. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

