Prediction of novel precursor miRNAs using a context-sensitive hidden Markov model (CSHMM)
- PMID: 20122201
- PMCID: PMC3009500
- DOI: 10.1186/1471-2105-11-S1-S29
Prediction of novel precursor miRNAs using a context-sensitive hidden Markov model (CSHMM)
Abstract
Background: It has been apparent in the last few years that small non coding RNAs (ncRNA) play a very significant role in biological regulation. Among these microRNAs (miRNAs), 22-23 nucleotide small regulatory RNAs, have been a major object of study as these have been found to be involved in some basic biological processes. So far about 706 miRNAs have been identified in humans alone. However, it is expected that there may be many more miRNAs encoded in the human genome. In this report, a "context-sensitive" Hidden Markov Model (CSHMM) to represent miRNA structures has been proposed and tested extensively. We also demonstrate how this model can be used in conjunction with filters as an ab initio method for miRNA identification.
Results: The probabilities of the CSHMM model were estimated using known human miRNA sequences. A classifier for miRNAs based on the likelihood score of this "trained" CSHMM was evaluated by: (a) cross-validation estimates using known human sequences, (b) predictions on a dataset of known miRNAs, and (c) prediction on a dataset of non coding RNAs. The CSHMM is compared with two recently developed methods, miPred and CID-miRNA. The results suggest that the CSHMM performs better than these methods. In addition, the CSHMM was used in a pipeline that includes filters that check for the presence of EST matches and the presence of Drosha cutting sites. This pipeline was used to scan and identify potential miRNAs from the human chromosome 19. It was also used to identify novel miRNAs from small RNA sequences of human normal leukocytes obtained by the Deep sequencing (Solexa) methodology. A total of 49 and 308 novel miRNAs were predicted from chromosome 19 and from the small RNA sequences respectively.
Conclusion: The results suggest that the CSHMM is likely to be a useful tool for miRNA discovery either for analysis of individual sequences or for genome scan. Our pipeline, consisting of a CSHMM and filters to reduce false positives shows promise as an approach for ab initio identification of novel miRNAs.
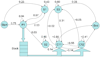
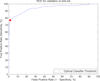
Figures
Similar articles
-
Ab initio identification of human microRNAs based on structure motifs.BMC Bioinformatics. 2007 Dec 18;8:478. doi: 10.1186/1471-2105-8-478. BMC Bioinformatics. 2007. PMID: 18088431 Free PMC article.
-
Identification of clustered microRNAs using an ab initio prediction method.BMC Bioinformatics. 2005 Nov 7;6:267. doi: 10.1186/1471-2105-6-267. BMC Bioinformatics. 2005. PMID: 16274478 Free PMC article.
-
Ab initio human miRNA and pre-miRNA prediction.J Bioinform Comput Biol. 2013 Dec;11(6):1343009. doi: 10.1142/S0219720013430099. Epub 2013 Dec 11. J Bioinform Comput Biol. 2013. PMID: 24372038
-
Analysis of microRNA transcriptome by deep sequencing of small RNA libraries of peripheral blood.BMC Genomics. 2010 May 7;11:288. doi: 10.1186/1471-2164-11-288. BMC Genomics. 2010. PMID: 20459673 Free PMC article.
-
Role of miRNA in carcinogenesis and biomarker selection: a methodological view.Expert Rev Mol Diagn. 2007 Sep;7(5):569-603. doi: 10.1586/14737159.7.5.569. Expert Rev Mol Diagn. 2007. PMID: 17892365 Review.
Cited by
-
miR-BAG: bagging based identification of microRNA precursors.PLoS One. 2012;7(9):e45782. doi: 10.1371/journal.pone.0045782. Epub 2012 Sep 25. PLoS One. 2012. PMID: 23049860 Free PMC article.
-
miRNAFold: a web server for fast miRNA precursor prediction in genomes.Nucleic Acids Res. 2016 Jul 8;44(W1):W181-4. doi: 10.1093/nar/gkw459. Epub 2016 May 29. Nucleic Acids Res. 2016. PMID: 27242364 Free PMC article.
-
CL-PMI: A Precursor MicroRNA Identification Method Based on Convolutional and Long Short-Term Memory Networks.Front Genet. 2019 Oct 11;10:967. doi: 10.3389/fgene.2019.00967. eCollection 2019. Front Genet. 2019. PMID: 31681416 Free PMC article.
-
Identification of real microRNA precursors with a pseudo structure status composition approach.PLoS One. 2015 Mar 30;10(3):e0121501. doi: 10.1371/journal.pone.0121501. eCollection 2015. PLoS One. 2015. PMID: 25821974 Free PMC article.
-
Mirinho: An efficient and general plant and animal pre-miRNA predictor for genomic and deep sequencing data.BMC Bioinformatics. 2015 May 29;16:179. doi: 10.1186/s12859-015-0594-0. BMC Bioinformatics. 2015. PMID: 26022464 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

