Conformation change in a self-recognizing autotransporter modulates bacterial cell-cell interaction
- PMID: 20123991
- PMCID: PMC2856270
- DOI: 10.1074/jbc.M109.069070
Conformation change in a self-recognizing autotransporter modulates bacterial cell-cell interaction
Abstract
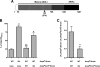
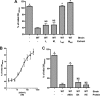
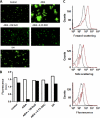
Bacteria mostly live as multicellular communities, although they are unicellular organisms, yet the mechanisms that tie individual bacteria together are often poorly understood. The adhesin involved in diffuse adherence (AIDA-I) is an adhesin of diarrheagenic Escherichia coli strains. AIDA-I also mediates bacterial auto-aggregation and biofilm formation and thus could be important for the organization of communities of pathogens. Using purified protein and whole bacteria, we provide direct evidence that AIDA-I promotes auto-aggregation by interacting with itself. Using various biophysical and biochemical techniques, we observed a conformational change in the protein during AIDA-AIDA interactions, strengthening the notion that this is a highly specific interaction. The self-association of AIDA-I is of high affinity but can be modulated by sodium chloride. We observe that a bile salt, sodium deoxycholate, also prevents AIDA-I oligomerization and bacterial auto-aggregation. Thus, we propose that AIDA-I, and most likely other similar autotransporters such as antigen 43 (Ag43) and TibA, organize bacterial communities of pathogens through a self-recognition mechanism that is sensitive to the environment. This could permit bacteria to switch between multicellular and unicellular lifestyles to complete their infection.
Figures
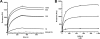

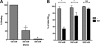
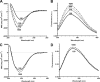
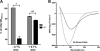
Similar articles
-
Glycosylation of the Escherichia coli TibA self-associating autotransporter influences the conformation and the functionality of the protein.PLoS One. 2013 Nov 20;8(11):e80739. doi: 10.1371/journal.pone.0080739. eCollection 2013. PLoS One. 2013. PMID: 24278316 Free PMC article.
-
O-linked glycosylation ensures the normal conformation of the autotransporter adhesin involved in diffuse adherence.J Bacteriol. 2007 Dec;189(24):8880-9. doi: 10.1128/JB.00969-07. Epub 2007 Oct 19. J Bacteriol. 2007. PMID: 17951390 Free PMC article.
-
Proteolytic processing is not essential for multiple functions of the Escherichia coli autotransporter adhesin involved in diffuse adherence (AIDA-I).J Bacteriol. 2006 Dec;188(24):8504-12. doi: 10.1128/JB.00864-06. Epub 2006 Oct 13. J Bacteriol. 2006. PMID: 17041044 Free PMC article.
-
Self-associating autotransporters, SAATs: functional and structural similarities.Int J Med Microbiol. 2006 Aug;296(4-5):187-95. doi: 10.1016/j.ijmm.2005.10.002. Int J Med Microbiol. 2006. PMID: 16600681 Review.
-
Autotransporter Adhesins in Escherichia coli Pathogenesis.Proteomics. 2017 Dec;17(23-24). doi: 10.1002/pmic.201600431. Epub 2017 Oct 12. Proteomics. 2017. PMID: 28665015 Review.
Cited by
-
Proteinaceous determinants of surface colonization in bacteria: bacterial adhesion and biofilm formation from a protein secretion perspective.Front Microbiol. 2013 Oct 14;4:303. doi: 10.3389/fmicb.2013.00303. Front Microbiol. 2013. PMID: 24133488 Free PMC article. Review.
-
Intra- and inter-species interactions within biofilms of important foodborne bacterial pathogens.Front Microbiol. 2015 Aug 20;6:841. doi: 10.3389/fmicb.2015.00841. eCollection 2015. Front Microbiol. 2015. PMID: 26347727 Free PMC article. Review.
-
Crystal structure of the Haemophilus influenzae Hap adhesin reveals an intercellular oligomerization mechanism for bacterial aggregation.EMBO J. 2011 Aug 12;30(18):3864-74. doi: 10.1038/emboj.2011.279. EMBO J. 2011. PMID: 21841773 Free PMC article.
-
Surface Plasmon Resonance: A Sensitive Tool to Study Protein-Protein Interactions.Methods Mol Biol. 2024;2715:363-382. doi: 10.1007/978-1-0716-3445-5_23. Methods Mol Biol. 2024. PMID: 37930540
-
Identification and mechanism of evolution of new alleles coding for the AIDA-I autotransporter of porcine pathogenic Escherichia coli.Appl Environ Microbiol. 2012 Jul;78(13):4597-605. doi: 10.1128/AEM.00906-12. Epub 2012 Apr 20. Appl Environ Microbiol. 2012. PMID: 22522689 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

