Combination of V106I and V179D polymorphic mutations in human immunodeficiency virus type 1 reverse transcriptase confers resistance to efavirenz and nevirapine but not etravirine
- PMID: 20124001
- PMCID: PMC2849364
- DOI: 10.1128/AAC.01480-09
Combination of V106I and V179D polymorphic mutations in human immunodeficiency virus type 1 reverse transcriptase confers resistance to efavirenz and nevirapine but not etravirine
Abstract
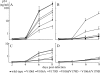
Etravirine (ETV) is a second-generation nonnucleoside reverse transcriptase (RT) inhibitor (NNRTI) introduced recently for salvage antiretroviral treatment after the emergence of NNRTI-resistant human immunodeficiency virus type 1 (HIV-1). Following its introduction, two naturally occurring mutations in HIV-1 RT, V106I and V179D, were listed as ETV resistance-associated mutations. However, the effect of these mutations on the development of NNRTI resistance has not been analyzed yet. To select highly NNRTI-resistant HIV-1 in vitro, monoclonal HIV-1 strains harboring V106I and V179D (HIV-1(V106I) and HIV-1(V179D)) were propagated in the presence of increasing concentrations of efavirenz (EFV). Interestingly, V179D emerged in one of three selection experiments from HIV-1(V106I) and V106I emerged in two of three experiments from HIV-1(V179D). Analysis of recombinant HIV-1 clones showed that the combination of V106I and V179D conferred significant resistance to EFV and nevirapine (NVP) but not to ETV. Structural analysis indicated that ETV can overcome the repulsive interactions caused by the combination of V106I and V179D through fine-tuning of its binding module to RT facilitated by its plastic structure, whereas EFV and NVP cannot because of their rigid structures. Analysis of clinical isolates showed comparable drug susceptibilities, and the same combination of mutations was found in some database patients who experienced virologic NNRTI-based treatment failure. The combination of V106I and V179D is a newly identified NNRTI resistance pattern of mutations. The combination of polymorphic and minor resistance-associated mutations should be interpreted carefully.
Figures


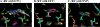
Similar articles
-
Constrained patterns of covariation and clustering of HIV-1 non-nucleoside reverse transcriptase inhibitor resistance mutations.J Antimicrob Chemother. 2010 Jul;65(7):1477-85. doi: 10.1093/jac/dkq140. Epub 2010 May 12. J Antimicrob Chemother. 2010. PMID: 20462946 Free PMC article.
-
Nonnucleoside reverse transcriptase inhibitor-resistant HIV is stimulated by efavirenz during early stages of infection.J Virol. 2011 Oct;85(20):10861-73. doi: 10.1128/JVI.05116-11. Epub 2011 Aug 10. J Virol. 2011. PMID: 21835788 Free PMC article.
-
Role of the K101E substitution in HIV-1 reverse transcriptase in resistance to rilpivirine and other nonnucleoside reverse transcriptase inhibitors.Antimicrob Agents Chemother. 2013 Nov;57(11):5649-57. doi: 10.1128/AAC.01536-13. Epub 2013 Sep 3. Antimicrob Agents Chemother. 2013. PMID: 24002090 Free PMC article.
-
HIV-1 reverse transcriptase and antiviral drug resistance. Part 2.Curr Opin Virol. 2013 Apr;3(2):119-28. doi: 10.1016/j.coviro.2013.03.014. Epub 2013 Apr 19. Curr Opin Virol. 2013. PMID: 23602470 Free PMC article. Review.
-
[Etravirine: genetic barrier and resistance development].Enferm Infecc Microbiol Clin. 2009 Dec;27 Suppl 2:32-9. doi: 10.1016/S0213-005X(09)73217-3. Enferm Infecc Microbiol Clin. 2009. PMID: 20116626 Review. Spanish.
Cited by
-
Distinct Clusters of HIV-1 CRF01_AE in Zhejiang, China: High-Risk Transmission Cluster 4 Requires Heightened Surveillance.Infect Drug Resist. 2024 Oct 11;17:4333-4342. doi: 10.2147/IDR.S480192. eCollection 2024. Infect Drug Resist. 2024. PMID: 39411499 Free PMC article.
-
In vitro HIV-1 evolution in response to triple reverse transcriptase inhibitors & in silico phenotypic analysis.PLoS One. 2013 Apr 17;8(4):e61102. doi: 10.1371/journal.pone.0061102. Print 2013. PLoS One. 2013. PMID: 23613794 Free PMC article.
-
The origin, dissemination, and molecular networks of HIV-1 CRF65_cpx strain in Hainan Island, China.BMC Infect Dis. 2024 Mar 1;24(1):269. doi: 10.1186/s12879-024-09101-w. BMC Infect Dis. 2024. PMID: 38424479 Free PMC article.
-
Evaluation of a Novel In-house HIV-1 Genotype Drug Resistance Assay using Clinical Samples in China.Curr HIV Res. 2022;20(1):32-41. doi: 10.2174/1570162X19666210910144433. Curr HIV Res. 2022. PMID: 34515004 Free PMC article.
-
Constrained patterns of covariation and clustering of HIV-1 non-nucleoside reverse transcriptase inhibitor resistance mutations.J Antimicrob Chemother. 2010 Jul;65(7):1477-85. doi: 10.1093/jac/dkq140. Epub 2010 May 12. J Antimicrob Chemother. 2010. PMID: 20462946 Free PMC article.
References
-
- Bacheler, L. T., E. D. Anton, P. Kudish, D. Baker, J. Bunville, K. Krakowski, L. Bolling, M. Aujay, X. V. Wang, D. Ellis, M. F. Becker, A. L. Lasut, H. J. George, D. R. Spalding, G. Hollis, and K. Abremski.2000. Human immunodeficiency virus type 1 mutations selected in patients failing efavirenz combination therapy. Antimicrob. Agents Chemother. 44:2475-2484. - PMC - PubMed
-
- Baxter, J. D., J. M. Schapiro, C. A. Boucher, V. M. Kohlbrenner, D. B. Hall, J. R. Scherer, and D. L. Mayers.2006. Genotypic changes in human immunodeficiency virus type 1 protease associated with reduced susceptibility and virologic response to the protease inhibitor tipranavir. J. Virol. 80:10794-10801. - PMC - PubMed
-
- Butler, I. F., I. Pandrea, P. A. Marx, and C. Apetrei.2007. HIV genetic diversity: biological and public health consequences. Curr. HIV Res. 5:23-45. - PubMed
-
- Das, K., A. D. Clark, Jr., P. J. Lewi, J. Heeres, M. R. De Jonge, L. M. Koymans, H. M. Vinkers, F. Daeyaert, D. W. Ludovici, M. J. Kukla, B. De Corte, R. W. Kavash, C. Y. Ho, H. He, M. A. Lichtenstein, K. Andries, R. Pauwels, M. P. De Bethune, P. L. Boyer, P. Clark, S. H. Hughes, P. A. Janssen, and E. Arnold.2004. Roles of conformational and positional adaptability in structure-based design of TMC125-R165335 (etravirine) and related non-nucleoside reverse transcriptase inhibitors that are highly potent and effective against wild-type and drug-resistant HIV-1 variants. J. Med. Chem. 47:2550-2560. - PubMed
-
- Duan, Y., C. Wu, S. Chowdhury, M. C. Lee, G. Xiong, W. Zhang, R. Yang, P. Cieplak, R. Luo, T. Lee, J. Caldwell, J. Wang, and P. Kollman.2003. A point-charged force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 24:1999-2012. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

