Distinct activities of exonuclease 1 and flap endonuclease 1 at telomeric g4 DNA
- PMID: 20126648
- PMCID: PMC2811187
- DOI: 10.1371/journal.pone.0008908
Distinct activities of exonuclease 1 and flap endonuclease 1 at telomeric g4 DNA
Abstract
Background: Exonuclease 1 (EXO1) and Flap endonuclease 1 (FEN1) are members of the RAD2 family of structure-specific nucleases. Genetic analysis has identified roles for EXO1 and FEN1 in replication, recombination, DNA repair and maintenance of telomeres. Telomeres are composed of G-rich repeats that readily form G4 DNA. We recently showed that human EXO1 and FEN1 exhibit distinct activities on G4 DNA substrates representative of intermediates in immunoglobulin class switch recombination.
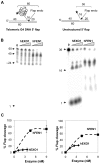
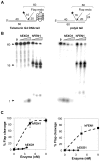
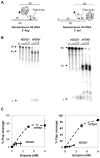
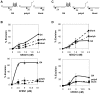
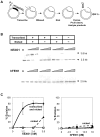
Methodology/principal findings: We have now compared activities of these enzymes on telomeric substrates bearing G4 DNA, identifying non-overlapping functions that provide mechanistic insight into the distinct telomeric phenotypes caused by their deficiencies. We show that hFEN1 but not hEXO1 cleaves substrates bearing telomeric G4 DNA 5'-flaps, consistent with the requirement for FEN1 in telomeric lagging strand replication. Both hEXO1 and hFEN1 are active on substrates bearing telomeric G4 DNA tails, resembling uncapped telomeres. Notably, hEXO1 but not hFEN1 is active on transcribed telomeric G-loops.
Conclusion/significance: Our results suggest that EXO1 may act at transcription-induced telomeric structures to promote telomere recombination while FEN1 has a dominant role in lagging strand replication at telomeres. Both enzymes can create ssDNA at uncapped telomere ends thereby contributing to recombination.
Conflict of interest statement
Figures

References
-
- Harrington JJ, Lieber MR. Functional domains within FEN-1 and RAD2 define a family of structure-specific endonucleases: implications for nucleotide excision repair. Genes Dev. 1994;8:1344–1355. - PubMed
-
- Bambara RA, Murante RS, Henricksen LA. Enzymes and reactions at the eukaryotic DNA replication fork. J Biol Chem. 1997;272:4647–4650. - PubMed
-
- Tran PT, Erdeniz N, Symington LS, Liskay RM. EXO1-A multi-tasking eukaryotic nuclease. DNA Repair (Amst) 2004;3:1549–1559. - PubMed
-
- Liu Y, Kao HI, Bambara RA. Flap endonuclease 1: a central component of DNA metabolism. Annu Rev Biochem. 2004;73:589–615. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

