The Arabidopsis wall associated kinase-like 10 gene encodes a functional guanylyl cyclase and is co-expressed with pathogen defense related genes
- PMID: 20126659
- PMCID: PMC2811198
- DOI: 10.1371/journal.pone.0008904
The Arabidopsis wall associated kinase-like 10 gene encodes a functional guanylyl cyclase and is co-expressed with pathogen defense related genes
Abstract
Background: Second messengers have a key role in linking environmental stimuli to physiological responses. One such messenger, guanosine 3',5'-cyclic monophosphate (cGMP), has long been known to be an essential signaling molecule in many different physiological processes in higher plants, including biotic stress responses. To date, however, the guanylyl cyclase (GC) enzymes that catalyze the formation of cGMP from GTP have largely remained elusive in higher plants.
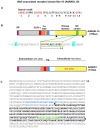
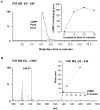
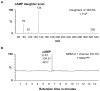
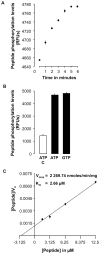
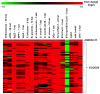
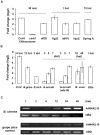
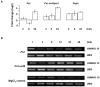
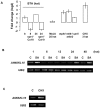
Principal findings: We have identified an Arabidopsis receptor type wall associated kinase-like molecule (AtWAKL10) as a candidate GC and provide experimental evidence to show that the intracellular domain of AtWAKL10(431-700) can generate cGMP in vitro. Further, we also demonstrate that the molecule has kinase activity indicating that AtWAKL10 is a twin-domain catalytic protein. A co-expression and stimulus-specific expression analysis revealed that AtWAKL10 is consistently co-expressed with well characterized pathogen defense related genes and along with these genes is induced early and sharply in response to a range of pathogens and their elicitors.
Conclusions: We demonstrate that AtWAKL10 is a twin-domain, kinase-GC signaling molecule that may function in biotic stress responses that are critically dependent on the second messenger cGMP.
Conflict of interest statement
Figures

Similar articles
-
The Arabidopsis thaliana proteome harbors undiscovered multi-domain molecules with functional guanylyl cyclase catalytic centers.Cell Commun Signal. 2013 Jul 8;11:48. doi: 10.1186/1478-811X-11-48. Cell Commun Signal. 2013. PMID: 23835195 Free PMC article.
-
Identification of a novel Arabidopsis thaliana nitric oxide-binding molecule with guanylate cyclase activity in vitro.FEBS Lett. 2011 Sep 2;585(17):2693-7. doi: 10.1016/j.febslet.2011.07.023. Epub 2011 Jul 31. FEBS Lett. 2011. PMID: 21820435
-
The Hippeastrum hybridum PepR1 gene (HpPepR1) encodes a functional guanylyl cyclase and is involved in early response to fungal infection.J Plant Physiol. 2017 Sep;216:100-107. doi: 10.1016/j.jplph.2017.05.024. Epub 2017 Jun 1. J Plant Physiol. 2017. PMID: 28609666
-
[Biosynthesis of cyclic GMP in plant cells - new insight into guanylate cyclases].Postepy Biochem. 2015;61(2):168-75. Postepy Biochem. 2015. PMID: 26689009 Review. Polish.
-
Regulation of photoreceptor membrane guanylyl cyclases by guanylyl cyclase activator proteins.Methods. 1999 Dec;19(4):521-31. doi: 10.1006/meth.1999.0894. Methods. 1999. PMID: 10581151 Review.
Cited by
-
Phaeophyceaean (Brown Algal) Extracts Activate Plant Defense Systems in Arabidopsis thaliana Challenged With Phytophthora cinnamomi.Front Plant Sci. 2020 Jul 7;11:852. doi: 10.3389/fpls.2020.00852. eCollection 2020. Front Plant Sci. 2020. PMID: 32765538 Free PMC article.
-
"Cyclic nucleotides in plants: from obscure messengers to central regulators".Front Plant Sci. 2025 Jun 25;16:1618243. doi: 10.3389/fpls.2025.1618243. eCollection 2025. Front Plant Sci. 2025. PMID: 40636015 Free PMC article. Review.
-
The Brassica napus wall-associated kinase-like (WAKL) gene Rlm9 provides race-specific blackleg resistance.Plant J. 2020 Nov;104(4):892-900. doi: 10.1111/tpj.14966. Epub 2020 Oct 7. Plant J. 2020. PMID: 32794614 Free PMC article.
-
Moonlighting Crypto-Enzymes and Domains as Ancient and Versatile Signaling Devices.Int J Mol Sci. 2024 Sep 2;25(17):9535. doi: 10.3390/ijms25179535. Int J Mol Sci. 2024. PMID: 39273482 Free PMC article. Review.
-
In Search of Enzymes with a Role in 3', 5'-Cyclic Guanosine Monophosphate Metabolism in Plants.Front Plant Sci. 2016 May 6;7:576. doi: 10.3389/fpls.2016.00576. eCollection 2016. Front Plant Sci. 2016. PMID: 27200049 Free PMC article. Review.
References
-
- Schaap P. Guanylyl cyclases across the tree of life. Front Biosci. 2005;10:1485–1498. - PubMed
-
- Pilz RB, Casteel DE. Regulation of gene expression by cyclic GMP. Circ Res. 2003;93:1034–1046. - PubMed
-
- Newton RP, Smith CJ. Cyclic nucleotides. Phytochemistry. 2004;65:2423–2437. - PubMed
-
- Prado AM, Porterfield DM, Feijo JA. Nitric oxide is involved in growth regulation and re-orientation of pollen tubes. Development. 2004;131:2707–2714. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

