Interaction of the exported malaria protein Pf332 with the red blood cell membrane skeleton
- PMID: 20132790
- PMCID: PMC4768738
- DOI: 10.1016/j.bbamem.2010.01.018
Interaction of the exported malaria protein Pf332 with the red blood cell membrane skeleton
Abstract
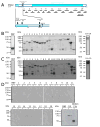
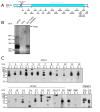
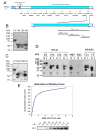
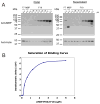
Intra-erythrocytic Plasmodium falciparum malaria parasites synthesize and export numerous proteins into the red blood cell (RBC) cytosol, where some bind to the RBC membrane skeleton. These interactions are responsible for the altered antigenic, morphological and functional properties of parasite-infected red blood cells (IRBCs). Plasmodium falciparum protein 332 (Pf332) is a large parasite protein that associates with the membrane skeleton and who's function has recently been elucidated. Using recombinant fragments of Pf332 in in vitro interaction assays, we have localised the specific domain within Pf332 that binds to the RBC membrane skeleton to an 86 residue sequence proximal to the C-terminus of Pf332. We have shown that this region partakes in a specific and saturable interaction with actin (K(d)=0.60 microM) but has no detectable affinity for spectrin. The only exported malaria protein previously known to bind to actin is PfEMP3 but here we demonstrate that there is no competition for actin-binding between PfEMP3 and Pf332, suggesting that they bind to different target sequences in actin.
2010 Elsevier B.V. All rights reserved.
Figures
Similar articles
-
Interactions between Plasmodium falciparum skeleton-binding protein 1 and the membrane skeleton of malaria-infected red blood cells.Biochim Biophys Acta. 2015 Jul;1848(7):1619-1628. doi: 10.1016/j.bbamem.2015.03.038. Epub 2015 Apr 14. Biochim Biophys Acta. 2015. PMID: 25883090 Free PMC article.
-
Tryptophan-rich domains of Plasmodium falciparum SURFIN4.2 and Plasmodium vivax PvSTP2 interact with membrane skeleton of red blood cell.Malar J. 2017 Mar 20;16(1):121. doi: 10.1186/s12936-017-1772-5. Malar J. 2017. PMID: 28320404 Free PMC article.
-
Plasmodium falciparum antigen 332 is a resident peripheral membrane protein of Maurer's clefts.PLoS One. 2012;7(11):e46980. doi: 10.1371/journal.pone.0046980. Epub 2012 Nov 20. PLoS One. 2012. PMID: 23185236 Free PMC article.
-
Vesicle-mediated trafficking of parasite proteins to the host cell cytosol and erythrocyte surface membrane in Plasmodium falciparum infected erythrocytes.Int J Parasitol. 2001 Oct;31(12):1381-91. doi: 10.1016/s0020-7519(01)00256-9. Int J Parasitol. 2001. PMID: 11566305 Review.
-
Feisty filaments: actin dynamics in the red blood cell membrane skeleton.Curr Opin Hematol. 2016 May;23(3):206-14. doi: 10.1097/MOH.0000000000000227. Curr Opin Hematol. 2016. PMID: 27055045 Free PMC article. Review.
Cited by
-
Plasmodium species: master renovators of their host cells.Nat Rev Microbiol. 2016 Aug;14(8):494-507. doi: 10.1038/nrmicro.2016.79. Epub 2016 Jul 4. Nat Rev Microbiol. 2016. PMID: 27374802 Review.
-
Host cell remodeling by pathogens: the exomembrane system in Plasmodium-infected erythrocytes.FEMS Microbiol Rev. 2016 Sep;40(5):701-21. doi: 10.1093/femsre/fuw016. FEMS Microbiol Rev. 2016. PMID: 27587718 Free PMC article. Review.
-
SURGE complex of Plasmodium falciparum in the rhoptry-neck (SURFIN4.2-RON4-GLURP) contributes to merozoite invasion.PLoS One. 2018 Aug 9;13(8):e0201669. doi: 10.1371/journal.pone.0201669. eCollection 2018. PLoS One. 2018. PMID: 30092030 Free PMC article.
-
An erythrocyte cytoskeleton-binding motif in exported Plasmodium falciparum proteins.Eukaryot Cell. 2011 Nov;10(11):1439-47. doi: 10.1128/EC.05180-11. Epub 2011 Sep 9. Eukaryot Cell. 2011. PMID: 21908595 Free PMC article.
-
Transposon mutagenesis identifies genes essential for Plasmodium falciparum gametocytogenesis.Proc Natl Acad Sci U S A. 2013 Apr 30;110(18):E1676-84. doi: 10.1073/pnas.1217712110. Epub 2013 Apr 9. Proc Natl Acad Sci U S A. 2013. PMID: 23572579 Free PMC article.
References
-
- Cooke BM, Mohandas N, Coppel RL. Malaria and the red blood cell membrane. Semin Hematol. 2004;41:173–188. - PubMed
-
- Walensky L, Narla M, Lux S., IV . Disorders of the Red Blood Cell Membrane. In: Handin R, Lux S IV, Stossel TP, editors. Blood: Princciples and Practice of Hematology. Lippincott, William and Wilkins; Philadelphia: 2003. pp. 1709–1858.
-
- Gascard P, Mohandas N. New insights into functions of erythroid proteins in nonerythroid cells. Curr Opin Hematol. 2000;7:123–129. - PubMed
-
- Karinch AM, Zimmer WE, Goodman SR. The Identification and Sequence of the Actin-binding Domain of Human Red Blood Cell β-Spectrin. J Biol Chem. 1990;265:11833–11840. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

