Dynamics of intracellular bacterial replication at the single cell level
- PMID: 20133586
- PMCID: PMC2840444
- DOI: 10.1073/pnas.1000041107
Dynamics of intracellular bacterial replication at the single cell level
Abstract
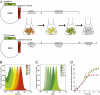
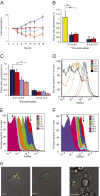
Several important pathogens cause disease by surviving and replicating within host cells. Bacterial proliferation is the product of both replication and killing undergone by the population. However, these processes are difficult to distinguish, and are usually assessed together by determination of net bacterial load. In addition, measurement of net load does not reveal heterogeneity within pathogen populations. This is particularly important in persistent infections in which slow or nongrowing bacteria are thought to have a major impact. Here we report the development of a reporter system based on fluorescence dilution that enables direct quantification of the replication dynamics of Salmonella enterica serovar Typhimurium (S. Typhimurium) in murine macrophages at both the population and single-cell level. We used this technique to demonstrate that a major S. Typhimurium virulence determinant, the Salmonella pathogenicity island 2 type III secretion system, is required for bacterial replication but does not have a major influence on resistance to killing. Furthermore, we found that, upon entry into macrophages, many bacteria do not replicate, but appear to enter a dormant-like state. These could represent an important reservoir of persistent bacteria. The approach could be extended to other pathogens to study the contribution of virulence and host resistance factors to replication and killing, and to identify and characterize nonreplicating bacteria associated with chronic or latent infections.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

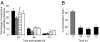
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

