Antigenic characterization of recombinant hemagglutinin proteins derived from different avian influenza virus subtypes
- PMID: 20140098
- PMCID: PMC2816723
- DOI: 10.1371/journal.pone.0009097
Antigenic characterization of recombinant hemagglutinin proteins derived from different avian influenza virus subtypes
Abstract
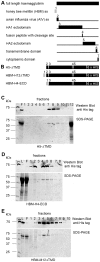
Since the advent of highly pathogenic variants of avian influenza virus (HPAIV), the main focus of avian influenza research has been the characterization and detection of HPAIV hemagglutinin (HA) from H5 and H7 subtypes. However, due to the high mutation and reassortation rate of influenza viruses, in theory any influenza strain may acquire increased pathogenicity irrespective of its subtype. A comprehensive antigenic characterization of influenza viruses encompassing all 16 HA and 9 neuraminidase subtypes will provide information useful for the design of differential diagnostic tools, and possibly, vaccines. We have expressed recombinant HA proteins from 3 different influenza virus HA subtypes in the baculovirus system. These proteins were used to generate polyclonal rabbit antisera, which were subsequently employed in epitope scanning analysis using peptide libraries spanning the entire HA. Here, we report the identification and characterization of linear, HA subtype-specific as well as inter subtype-conserved epitopes along the HA proteins. Selected subtype-specific epitopes were shown to be suitable for the differentiation of anti-HA antibodies in an ELISA.
Conflict of interest statement
Figures

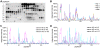


Similar articles
-
A novel epitope-blocking ELISA for specific and sensitive detection of antibodies against H5-subtype influenza virus hemagglutinin.Virol J. 2021 Apr 30;18(1):91. doi: 10.1186/s12985-021-01564-6. Virol J. 2021. PMID: 33931074 Free PMC article.
-
Cross-protective potential of a novel monoclonal antibody directed against antigenic site B of the hemagglutinin of influenza A viruses.PLoS Pathog. 2009 Mar;5(3):e1000350. doi: 10.1371/journal.ppat.1000350. Epub 2009 Mar 20. PLoS Pathog. 2009. PMID: 19300497 Free PMC article.
-
Amino acid substitutions in antigenic region B of hemagglutinin play a critical role in the antigenic drift of subclade 2.3.4.4 highly pathogenic H5NX influenza viruses.Transbound Emerg Dis. 2020 Jan;67(1):263-275. doi: 10.1111/tbed.13347. Epub 2019 Sep 28. Transbound Emerg Dis. 2020. PMID: 31484213
-
Epitopes in the HA and NA of H5 and H7 avian influenza viruses that are important for antigenic drift.FEMS Microbiol Rev. 2024 May 8;48(3):fuae014. doi: 10.1093/femsre/fuae014. FEMS Microbiol Rev. 2024. PMID: 38734891 Free PMC article. Review.
-
The antigenic architecture of the hemagglutinin of influenza H5N1 viruses.Mol Immunol. 2013 Dec;56(4):705-19. doi: 10.1016/j.molimm.2013.07.010. Epub 2013 Aug 7. Mol Immunol. 2013. PMID: 23933511 Review.
Cited by
-
Serological characterization of guinea pigs infected with H3N2 human influenza or immunized with hemagglutinin protein.Virol J. 2010 Aug 24;7:200. doi: 10.1186/1743-422X-7-200. Virol J. 2010. PMID: 20735849 Free PMC article.
-
DNA Vaccine Encoding a Modified Hemagglutinin Trimer of Avian Influenza A Virus H5N8 Protects Mice from Viral Challenge.Vaccines (Basel). 2024 May 14;12(5):538. doi: 10.3390/vaccines12050538. Vaccines (Basel). 2024. PMID: 38793789 Free PMC article.
-
Prediction of B-cell linear epitopes with a combination of support vector machine classification and amino acid propensity identification.J Biomed Biotechnol. 2011;2011:432830. doi: 10.1155/2011/432830. Epub 2011 Aug 23. J Biomed Biotechnol. 2011. PMID: 21876642 Free PMC article.
-
Insight into highly conserved H1 subtype-specific epitopes in influenza virus hemagglutinin.PLoS One. 2014 Feb 26;9(2):e89803. doi: 10.1371/journal.pone.0089803. eCollection 2014. PLoS One. 2014. PMID: 24587046 Free PMC article.
-
Heterosubtypic immunity to influenza A virus infections in mallards may explain existence of multiple virus subtypes.PLoS Pathog. 2013;9(6):e1003443. doi: 10.1371/journal.ppat.1003443. Epub 2013 Jun 20. PLoS Pathog. 2013. PMID: 23818849 Free PMC article.
References
-
- Suzuki Y. Sialobiology of influenza: molecular mechanism of host range variation of influenza viruses. Biol Pharm Bull. 2005;28:399–408. - PubMed
-
- Kalthoff D, Globig A, Beer M. (Highly pathogenic) avian influenza as a zoonotic agent. Vet Microbiol 2009 - PubMed
-
- Kelly TR, Hawkins MG, Sandrock CE, Boyce WM. A review of highly pathogenic avian influenza in birds, with an emphasis on Asian H5N1 and recommendations for prevention and control. J Avian Med Surg. 2008;22:1–16. - PubMed
-
- OIE. Manual of Diagnostic Tests and Vaccines for Terrestrial Animals. ISBN. 2008. 978-92-9044-718-4. Vol. 1. Chapter 2.3.4.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

