Genomic hotspots for adaptation: the population genetics of Müllerian mimicry in the Heliconius melpomene clade
- PMID: 20140188
- PMCID: PMC2816687
- DOI: 10.1371/journal.pgen.1000794
Genomic hotspots for adaptation: the population genetics of Müllerian mimicry in the Heliconius melpomene clade
Abstract
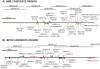
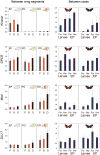
Wing patterning in Heliconius butterflies is a longstanding example of both Müllerian mimicry and phenotypic radiation under strong natural selection. The loci controlling such patterns are "hotspots" for adaptive evolution with great allelic diversity across different species in the genus. We characterise nucleotide variation, genotype-by-phenotype associations, linkage disequilibrium, and candidate gene expression at two loci and across multiple hybrid zones in Heliconius melpomene and relatives. Alleles at HmB control the presence or absence of the red forewing band, while alleles at HmYb control the yellow hindwing bar. Across HmYb two regions, separated by approximately 100 kb, show significant genotype-by-phenotype associations that are replicated across independent hybrid zones. In contrast, at HmB a single peak of association indicates the likely position of functional sites at three genes, encoding a kinesin, a G-protein coupled receptor, and an mRNA splicing factor. At both HmYb and HmB there is evidence for enhanced linkage disequilibrium (LD) between associated sites separated by up to 14 kb, suggesting that multiple sites are under selection. However, there was no evidence for reduced variation or deviations from neutrality that might indicate a recent selective sweep, consistent with these alleles being relatively old. Of the three genes showing an association with the HmB locus, the kinesin shows differences in wing disc expression between races that are replicated in the co-mimic, Heliconius erato, providing striking evidence for parallel changes in gene expression between Müllerian co-mimics. Wing patterning loci in Heliconius melpomene therefore show a haplotype structure maintained by selection, but no evidence for a recent selective sweep. The complex genetic pattern contrasts with the simple genetic basis of many adaptive traits studied previously, but may provide a better model for most adaptation in natural populations that has arisen over millions rather than tens of years.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Onuma Y, Takahashi S, Asashima M, Kurata S, Gehring WJ. Conservation of Pax 6 function and upstream activation by Notch signaling in eye development of frogs and flies. Proceedings of the National Academy of Sciences of the United States of America. 2002;99:2020–2025. doi: 10.1073/pnas.022626999. - DOI - PMC - PubMed
-
- Mundy NI. A window on the genetics of evolution: MC1R and plumage colouration in birds. Proceedings of the Royal Society B: Biological Sciences. 2005;272:1633–1640. doi: 10.1098/rspb.2005.3107. - DOI - PMC - PubMed
-
- Hoekstra HE, Hirschmann RJ, Bundey RA, Insel PA, Crossland JP. A single amino acid mutation contributes to adaptive beach mouse color pattern. Science. 2006;313:101–4. doi: 10.1126/science.1126121. - DOI - PubMed
-
- Nachman MW, Hoekstra HE, D'Agostino SL. The genetic basis of adaptive melanism in pocket mice. Proc Natl Acad Sci U S A. 2003;100:5268–5273. doi: 10.1073/pnas.0431157100. - DOI - PMC - PubMed
-
- Colosimo PF, Hosemann KE, Balabhadra S, Villarreal G, Dickson M, et al. Widespread Parallel Evolution in Sticklebacks by Repeated Fixation of Ectodysplasin Alleles. Science. 2005;307:1928–1933. doi: 10.1126/science.1107239. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

