Structure and kinetic investigation of Streptococcus pyogenes family GH38 alpha-mannosidase
- PMID: 20140249
- PMCID: PMC2815779
- DOI: 10.1371/journal.pone.0009006
Structure and kinetic investigation of Streptococcus pyogenes family GH38 alpha-mannosidase
Abstract
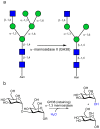
Background: The enzymatic hydrolysis of alpha-mannosides is catalyzed by glycoside hydrolases (GH), termed alpha-mannosidases. These enzymes are found in different GH sequence-based families. Considerable research has probed the role of higher eukaryotic "GH38" alpha-mannosides that play a key role in the modification and diversification of hybrid N-glycans; processes with strong cellular links to cancer and autoimmune disease. The most extensively studied of these enzymes is the Drosophila GH38 alpha-mannosidase II, which has been shown to be a retaining alpha-mannosidase that targets both alpha-1,3 and alpha-1,6 mannosyl linkages, an activity that enables the enzyme to process GlcNAc(Man)(5)(GlcNAc)(2) hybrid N-glycans to GlcNAc(Man)(3)(GlcNAc)(2). Far less well understood is the observation that many bacterial species, predominantly but not exclusively pathogens and symbionts, also possess putative GH38 alpha-mannosidases whose activity and specificity is unknown.
Methodology/principal findings: Here we show that the Streptococcus pyogenes (M1 GAS SF370) GH38 enzyme (Spy1604; hereafter SpGH38) is an alpha-mannosidase with specificity for alpha-1,3 mannosidic linkages. The 3D X-ray structure of SpGH38, obtained in native form at 1.9 A resolution and in complex with the inhibitor swainsonine (K(i) 18 microM) at 2.6 A, reveals a canonical GH38 five-domain structure in which the catalytic "-1" subsite shows high similarity with the Drosophila enzyme, including the catalytic Zn(2+) ion. In contrast, the "leaving group" subsites of SpGH38 display considerable differences to the higher eukaryotic GH38s; features that contribute to their apparent specificity.
Conclusions/significance: Although the in vivo function of this streptococcal GH38 alpha-mannosidase remains unknown, it is shown to be an alpha-mannosidase active on N-glycans. SpGH38 lies on an operon that also contains the GH84 hexosaminidase (Spy1600) and an additional putative glycosidase. The activity of SpGH38, together with its genomic context, strongly hints at a function in the degradation of host N- or possibly O-glycans. The absence of any classical signal peptide further suggests that SpGH38 may be intracellular, perhaps functioning in the subsequent degradation of extracellular host glycans following their initial digestion by secreted glycosidases.
Conflict of interest statement
Figures
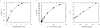
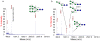
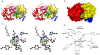
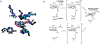

References
-
- Davis BG. Hand in Glove - Investigating Glycocode. Chem Ind. 2000:134–138.
-
- Ito Y, Ogawa T. A novel approach to the stereoselective synthesis of beta-mannosides. Angew Chemie Int Ed. 1994;33:1765–1767.
-
- Crich D, Chandrasekera NS. Mechanism of 4,6-0-benzylidene-directed beta-Mannosylation as determined by alpha-deuterium kinetic isotope effects. Angew Chemie Int Ed. 2004;43:5386–5389. - PubMed
-
- Gridley JJ, Osborn HMI. Recent advances in the construction of beta-D-mannose and beta-D-mannosamine linkages. J Chem Soc Perkin Trans 1. 2000;10:1471–1491.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

