Biosynthesis of rhizocticins, antifungal phosphonate oligopeptides produced by Bacillus subtilis ATCC6633
- PMID: 20142038
- PMCID: PMC2819989
- DOI: 10.1016/j.chembiol.2009.11.017
Biosynthesis of rhizocticins, antifungal phosphonate oligopeptides produced by Bacillus subtilis ATCC6633
Abstract
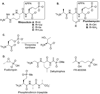
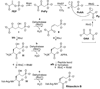
Rhizocticins are phosphonate oligopeptide antibiotics containing the C-terminal nonproteinogenic amino acid (Z)-l-2-amino-5-phosphono-3-pentenoic acid (APPA). Here we report the identification and characterization of the rhizocticin biosynthetic gene cluster (rhi) in Bacillus subtilis ATCC6633. Rhizocticin B was heterologously produced in the nonproducer strain Bacillus subtilis 168. A biosynthetic pathway is proposed on the basis of bioinformatics analysis of the rhi genes. One of the steps during the biosynthesis of APPA is an unusual aldol reaction between phosphonoacetaldehyde and oxaloacetate catalyzed by an aldolase homolog RhiG. Recombinant RhiG was prepared, and the product of an in vitro enzymatic conversion was characterized. Access to this intermediate allows for biochemical characterization of subsequent steps in the pathway.
Copyright (c) 2010 Elsevier Ltd. All rights reserved.
Figures



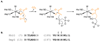
References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

