Distinct, ecotype-specific genome and proteome signatures in the marine cyanobacteria Prochlorococcus
- PMID: 20146791
- PMCID: PMC2836286
- DOI: 10.1186/1471-2164-11-103
Distinct, ecotype-specific genome and proteome signatures in the marine cyanobacteria Prochlorococcus
Abstract
Background: The marine cyanobacterium Prochlorococcus marinus, having multiple ecotypes of distinct genotypic/phenotypic traits and being the first documented example of genome shrinkage in free-living organisms, offers an ideal system for studying niche-driven molecular micro-diversity in closely related microbes. The present study, through an extensive comparative analysis of various genomic/proteomic features of 6 high light (HL) and 6 low light (LL) adapted strains, makes an attempt to identify molecular determinants associated with their vertical niche partitioning.
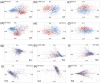
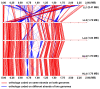
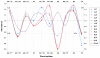
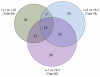
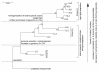
Results: Pronounced strand-specific asymmetry in synonymous codon usage is observed exclusively in LL strains. Distinct dinucleotide abundance profiles are exhibited by 2 LL strains with larger genomes and G+C-content approximately 50% (group LLa), 4 LL strains having reduced genomes and G+C-content approximately 35-37% (group LLb), and 6 HL strains. Taking into account the emergence of LLa, LLb and HL strains (based on 16S rRNA phylogeny), a gradual increase in average aromaticity, pI values and beta- & coil-forming propensities and a decrease in mean hydrophobicity, instability indices and helix-forming propensities of core proteins are observed. Greater variations in orthologous gene repertoire are found between LLa and LLb strains, while higher number of positively selected genes exist between LL and HL strains.
Conclusion: Strains of different Prochlorococcus groups are characterized by distinct compositional, physicochemical and structural traits that are not mere remnants of a continuous genetic drift, but are potential outcomes of a grand scheme of niche-oriented stepwise diversification, that might have driven them chronologically towards greater stability/fidelity and invoked upon them a special ability to inhabit diverse oceanic environments.
Figures

Similar articles
-
Whole genome phylogeny of Prochlorococcus marinus group of cyanobacteria: genome alignment and overlapping gene approach.Interdiscip Sci. 2014 Jun;6(2):149-57. doi: 10.1007/s12539-013-0024-9. Epub 2014 Jun 17. Interdiscip Sci. 2014. PMID: 25172453
-
Novel isolates expand the physiological diversity of Prochlorococcus and illuminate its macroevolution.mBio. 2024 Nov 13;15(11):e0349723. doi: 10.1128/mbio.03497-23. Epub 2024 Oct 18. mBio. 2024. PMID: 39422514 Free PMC article.
-
Patterns and implications of gene gain and loss in the evolution of Prochlorococcus.PLoS Genet. 2007 Dec;3(12):e231. doi: 10.1371/journal.pgen.0030231. PLoS Genet. 2007. PMID: 18159947 Free PMC article.
-
Ecological genomics of marine picocyanobacteria.Microbiol Mol Biol Rev. 2009 Jun;73(2):249-99. doi: 10.1128/MMBR.00035-08. Microbiol Mol Biol Rev. 2009. PMID: 19487728 Free PMC article. Review.
-
Phycobiliproteins in Prochlorococcus marinus: biosynthesis of pigments and their assembly into proteins.Eur J Cell Biol. 2010 Dec;89(12):1005-10. doi: 10.1016/j.ejcb.2010.06.017. Epub 2010 Aug 17. Eur J Cell Biol. 2010. PMID: 20724022 Review.
Cited by
-
Divergences in gene repertoire among the reference Prevotella genomes derived from distinct body sites of human.BMC Genomics. 2015 Mar 5;16(1):153. doi: 10.1186/s12864-015-1350-6. BMC Genomics. 2015. PMID: 25887946 Free PMC article.
-
A tribute to disorder in the genome of the bloom-forming freshwater cyanobacterium Microcystis aeruginosa.PLoS One. 2013 Aug 12;8(8):e70747. doi: 10.1371/journal.pone.0070747. eCollection 2013. PLoS One. 2013. PMID: 23950996 Free PMC article.
-
Excess of non-conservative amino acid changes in marine bacterioplankton lineages with reduced genomes.Nat Microbiol. 2017 Jun 12;2:17091. doi: 10.1038/nmicrobiol.2017.91. Nat Microbiol. 2017. PMID: 28604700
-
Toward a systems-level understanding of gene regulatory, protein interaction, and metabolic networks in cyanobacteria.Front Genet. 2014 Jul 2;5:191. doi: 10.3389/fgene.2014.00191. eCollection 2014. Front Genet. 2014. PMID: 25071821 Free PMC article. Review.
-
Mutational pressure dictates synonymous codon usage in freshwater unicellular α - cyanobacterial descendant Paulinella chromatophora and β - cyanobacterium Synechococcus elongatus PCC6301.Springerplus. 2013 Sep 30;2:492. doi: 10.1186/2193-1801-2-492. eCollection 2013. Springerplus. 2013. PMID: 24255825 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

