Caulobacter PopZ forms a polar subdomain dictating sequential changes in pole composition and function
- PMID: 20149103
- PMCID: PMC2935252
- DOI: 10.1111/j.1365-2958.2010.07088.x
Caulobacter PopZ forms a polar subdomain dictating sequential changes in pole composition and function
Abstract
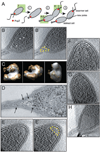
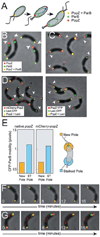
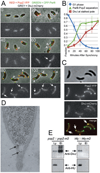
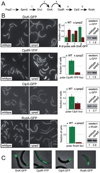
The bacterium Caulobacter crescentus has morphologically and functionally distinct cell poles that undergo sequential changes during the cell cycle. We show that the PopZ oligomeric network forms polar ribosome exclusion zones that change function during cell cycle progression. The parS/ParB chromosomal centromere is tethered to PopZ at one pole prior to the initiation of DNA replication. During polar maturation, the PopZ-centromere tether is broken, and the PopZ zone at that pole then switches function to act as a recruitment factor for the ordered addition of multiple proteins that promote the transformation of the flagellated pole into a stalked pole. Stalked pole assembly, in turn, triggers the initiation of chromosome replication, which signals the formation of a new PopZ zone at the opposite cell pole, where it functions to anchor the newly duplicated centromere that has traversed the long axis of the cell. We propose that pole-specific control of PopZ function co-ordinates polar development and cell cycle progression by enabling independent assembly and tethering activities at the two cell poles.
Figures


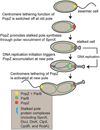
References
-
- Aaron M, Charbon G, Lam H, Schwarz H, Vollmer W, Jacobs-Wagner C. The tubulin homologue FtsZ contributes to cell elongation by guiding cell wall precursor synthesis in Caulobacter crescentus. Mol Microbiol. 2007;64:938–952. - PubMed
-
- Baker TA, Funnell BE, Kornberg A. Helicase action of dnaB protein during replication from the Escherichia coli chromosomal origin in vitro. J Biol Chem. 1987;262:6877–6885. - PubMed
-
- Bowers LM, Shapland EB, Ryan KR. Who’s in charge here? Regulating cell cycle regulators. Curr Opin Microbiol. 2008;11:547–552. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

