Neuronal regulation of the spatial patterning of neurogenesis
- PMID: 20152184
- PMCID: PMC2822724
- DOI: 10.1016/j.devcel.2009.11.010
Neuronal regulation of the spatial patterning of neurogenesis
Abstract
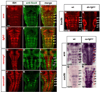
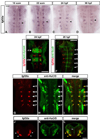
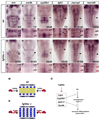
Precise regulation of neurogenesis is achieved in specific regions of the vertebrate nervous system by formation of distinct neurogenic and nonneurogenic zones. We have investigated how neurogenesis becomes confined to zones adjacent to rhombomere boundaries in the zebrafish hindbrain. The nonneurogenic zone at segment centers comprises a distinct progenitor population that expresses fibroblast growth factor (fgfr) 2, erm, sox9b, and the retinoic acid degrading enzyme, cyp26b1. FGF receptor activation upregulates expression of these genes and inhibits neurogenesis in segment centers. Cyp26 activity is a key effector inhibiting neuronal differentiation, suggesting antagonistic interactions with retinoid signaling. We identify the critical FGF ligand, fgf20a, which is expressed by specific neurons located in the mantle region at the center of segments, adjacent to the nonneurogenic zone. Fgf20a mutants have ectopic neurogenesis and lack the segment center progenitor population. Our findings reveal how signaling from neurons induces formation of a nonneurogenic zone of neural progenitors.
(c) 2010 Elsevier Inc. All rights reserved.
Figures




Similar articles
-
Signalling from hindbrain boundaries regulates neuronal clustering that patterns neurogenesis.Development. 2012 Aug;139(16):2978-87. doi: 10.1242/dev.080135. Epub 2012 Jul 4. Development. 2012. PMID: 22764046 Free PMC article.
-
vhnf1 integrates global RA patterning and local FGF signals to direct posterior hindbrain development in zebrafish.Development. 2004 Sep;131(18):4511-20. doi: 10.1242/dev.01297. Development. 2004. PMID: 15342476
-
Opposing actions of histone deacetylase 1 and Notch signalling restrict expression of erm and fgf20a to hindbrain rhombomere centres during zebrafish neurogenesis.Int J Dev Biol. 2011;55(6):597-602. doi: 10.1387/ijdb.113315el. Int J Dev Biol. 2011. PMID: 21948707
-
ArRAnging the hindbrain.Trends Neurosci. 2002 Feb;25(2):61-4. doi: 10.1016/s0166-2236(02)02067-2. Trends Neurosci. 2002. PMID: 11814548 Review.
-
How degrading: Cyp26s in hindbrain development.Dev Dyn. 2008 Oct;237(10):2775-90. doi: 10.1002/dvdy.21695. Dev Dyn. 2008. PMID: 18816852 Free PMC article. Review.
Cited by
-
Hindbrain boundaries as niches of neural progenitor and stem cells regulated by the extracellular matrix proteoglycan chondroitin sulphate.Development. 2024 Feb 15;151(4):dev201934. doi: 10.1242/dev.201934. Epub 2024 Feb 13. Development. 2024. PMID: 38251863 Free PMC article.
-
Temporal changes of cytochrome P450 (Cyp) and eicosanoid-related gene expression in the rat brain after traumatic brain injury.BMC Genomics. 2013 May 4;14:303. doi: 10.1186/1471-2164-14-303. BMC Genomics. 2013. PMID: 23642095 Free PMC article.
-
Forced to communicate: Integration of mechanical and biochemical signaling in morphogenesis.Curr Opin Cell Biol. 2020 Oct;66:59-68. doi: 10.1016/j.ceb.2020.05.004. Epub 2020 Jun 20. Curr Opin Cell Biol. 2020. PMID: 32569947 Free PMC article. Review.
-
FGF signalling restricts haematopoietic stem cell specification via modulation of the BMP pathway.Nat Commun. 2014 Nov 27;5:5588. doi: 10.1038/ncomms6588. Nat Commun. 2014. PMID: 25429520 Free PMC article.
-
The Thalamus Regulates Retinoic Acid Signaling and Development of Parvalbumin Interneurons in Postnatal Mouse Prefrontal Cortex.eNeuro. 2019 Mar 11;6(1):ENEURO.0018-19.2019. doi: 10.1523/ENEURO.0018-19.2019. eCollection 2019 Jan-Feb. eNeuro. 2019. PMID: 30868103 Free PMC article.
References
-
- Amoyel M, Cheng YC, Jiang YJ, Wilkinson DG. Wnt1 regulates neurogenesis and mediates lateral inhibition of boundary cell specification in the zebrafish hindbrain. Development. 2005;132:775–785. - PubMed
-
- Bae YK, Shimizu T, Hibi M. Patterning of proneuronal and interproneuronal domains by hairy- and enhancer of split-related genes in zebrafish neuroectoderm. Development. 2005;132:1375–1385. - PubMed
-
- Bally-Cuif L, Hammerschmidt M. Induction and patterning of neuronal development, and its connection to cell cycle control. Curr Opin Neurobiol. 2003;13:16–25. - PubMed
-
- Bertrand N, Castro DS, Guillemot F. Proneural genes and the specification of neural cell types. Nat Rev Neurosci. 2002;3:517–530. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

