Isolation and characterization of selective and potent human Fab inhibitors directed to the active-site region of the two-component NS2B-NS3 proteinase of West Nile virus
- PMID: 20156198
- PMCID: PMC2958048
- DOI: 10.1042/BJ20100074
Isolation and characterization of selective and potent human Fab inhibitors directed to the active-site region of the two-component NS2B-NS3 proteinase of West Nile virus
Abstract
There is a need to develop inhibitors of mosquito-borne flaviviruses, including WNV (West Nile virus). In the present paper, we describe a novel and efficient recombinant-antibody technology that led us to the isolation of inhibitory high-affinity human antibodies to the active-site region of a viral proteinase. As a proof-of-principal, we have successfully used this technology and the synthetic naive human combinatorial antibody library HuCAL GOLD(R) to isolate selective and potent function-blocking active-site-targeting antibodies to the two-component WNV NS (non-structural protein) 2B-NS3 serine proteinase, the only proteinase encoded by the flaviviral genome. First, we used the wild-type enzyme in antibody screens. Next, the positive antibody clones were counter-screened using an NS2B-NS3 mutant with a single mutation of the catalytically essential active-site histidine residue. The specificity of the antibodies to the active site was confirmed by substrate-cleavage reactions and also by using proteinase mutants with additional single amino-acid substitutions in the active-site region. The selected WNV antibodies did not recognize the structurally similar viral proteinases from Dengue virus type 2 and hepatitis C virus, and human serine proteinases. Because of their high selectivity and affinity, the identified human antibodies are attractive reagents for both further mutagenesis and structure-based optimization and, in addition, for studies of NS2B-NS3 activity. Conceptually, it is likely that the generic technology reported in the present paper will be useful for the generation of active-site-specific antibody probes for multiple enzymes.
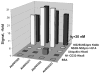
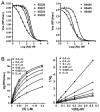
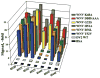
Figures


Similar articles
-
Structure-based mutagenesis identifies important novel determinants of the NS2B cofactor of the West Nile virus two-component NS2B-NS3 proteinase.J Gen Virol. 2008 Mar;89(Pt 3):636-641. doi: 10.1099/vir.0.83359-0. J Gen Virol. 2008. PMID: 18272753
-
Profiling of flaviviral NS2B-NS3 protease specificity provides a structural basis for the development of selective chemical tools that differentiate Dengue from Zika and West Nile viruses.Antiviral Res. 2020 Mar;175:104731. doi: 10.1016/j.antiviral.2020.104731. Epub 2020 Jan 31. Antiviral Res. 2020. PMID: 32014497
-
Homology modeling and molecular dynamics study of West Nile virus NS3 protease: a molecular basis for the catalytic activity increased by the NS2B cofactor.Proteins. 2006 Nov 15;65(3):692-701. doi: 10.1002/prot.21129. Proteins. 2006. PMID: 16972281
-
West Nile Virus NS2B/NS3 protease as an antiviral target.Curr Med Chem. 2008;15(27):2771-84. doi: 10.2174/092986708786242804. Curr Med Chem. 2008. PMID: 18991636 Review.
-
Potential Role of Flavivirus NS2B-NS3 Proteases in Viral Pathogenesis and Anti-flavivirus Drug Discovery Employing Animal Cells and Models: A Review.Viruses. 2021 Dec 28;14(1):44. doi: 10.3390/v14010044. Viruses. 2021. PMID: 35062249 Free PMC article. Review.
Cited by
-
Structural and functional parameters of the flaviviral protease: a promising antiviral drug target.Future Virol. 2010 Sep 1;5(5):593-606. doi: 10.2217/fvl.10.39. Future Virol. 2010. PMID: 21076642 Free PMC article.
-
High affinity human antibody fragments to dengue virus non-structural protein 3.PLoS Negl Trop Dis. 2010 Nov 9;4(11):e881. doi: 10.1371/journal.pntd.0000881. PLoS Negl Trop Dis. 2010. PMID: 21085466 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

