Widespread genomic instability mediated by a pathway involving glycoprotein Ib alpha and Aurora B kinase
- PMID: 20157117
- PMCID: PMC2857109
- DOI: 10.1074/jbc.M109.084913
Widespread genomic instability mediated by a pathway involving glycoprotein Ib alpha and Aurora B kinase
Abstract
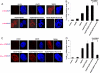
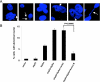
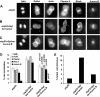
c-Myc (Myc) oncoprotein induction of genomic instability (GI) contributes to its initial transforming function and subsequent tumor cell evolution. We describe here a pathway by which Myc, via its target protein glycoprotein Ibalpha (GpIb alpha), mediates GI. Proteomic profiling revealed that the serine/threonine kinase Aurora B is down-regulated by GpIb alpha in p53-deficient primary human fibroblasts. The phenotypes of Aurora B deficiency are strikingly reminiscent of Myc or GpIb alpha overexpression and include double-stranded DNA breaks, altered nuclear size and morphology, chromatin bridges, cleavage furrow regression, and tetraploidy. During mitosis, GpIb alpha and Aurora B redistribute to the cleavage furrow along with other cleavage furrow proteins. GpIb alpha overexpression at levels comparable with those seen in some tumor cells causes the dispersal of these proteins but not Aurora B, resulting in furrow regression and cytokinesis failure. Aurora B normalization redirects the mislocalized furrow proteins to their proper location, corrects the cleavage furrow abnormalities, and restores genomic stability. Aurora B thus appears necessary for a previously unrecognized function in guiding and positioning a number of key proteins, including GpIb alpha to the cleavage furrow. These findings underscore the importance of maintaining a delicate balance among cleavage furrow-associated proteins during mitosis. Suppression of Aurora B via GpIb alpha provides a unifying and mechanistic explanation for several types of Myc-mediated GI.
Figures




Similar articles
-
Aurora-B and Rho-kinase/ROCK, the two cleavage furrow kinases, independently regulate the progression of cytokinesis: possible existence of a novel cleavage furrow kinase phosphorylates ezrin/radixin/moesin (ERM).Genes Cells. 2005 Feb;10(2):127-37. doi: 10.1111/j.1365-2443.2005.00824.x. Genes Cells. 2005. PMID: 15676024
-
Down-regulation of Aurora B kinase induces cellular senescence in human fibroblasts and endothelial cells through a p53-dependent pathway.FEBS Lett. 2011 Nov 16;585(22):3569-76. doi: 10.1016/j.febslet.2011.10.022. Epub 2011 Oct 20. FEBS Lett. 2011. PMID: 22024481
-
Therapeutic potential of a synthetic lethal interaction between the MYC proto-oncogene and inhibition of aurora-B kinase.Proc Natl Acad Sci U S A. 2010 Aug 3;107(31):13836-41. doi: 10.1073/pnas.1008366107. Epub 2010 Jul 19. Proc Natl Acad Sci U S A. 2010. PMID: 20643922 Free PMC article.
-
[Aurora kinases and cancer].Gan To Kagaku Ryoho. 2005 Jan;32(1):1-5. Gan To Kagaku Ryoho. 2005. PMID: 15675572 Review. Japanese.
-
Aurora B: a new prognostic marker and therapeutic target in cancer.Curr Med Chem. 2011;18(4):482-96. doi: 10.2174/092986711794480203. Curr Med Chem. 2011. PMID: 21143115 Review.
Cited by
-
Unbalanced replication as a major source of genetic instability in cancer cells.Am J Blood Res. 2012;2(3):160-9. Epub 2012 Oct 20. Am J Blood Res. 2012. PMID: 23119227 Free PMC article.
-
MicroRNAs, genomic instability and cancer.Int J Mol Sci. 2014 Aug 20;15(8):14475-91. doi: 10.3390/ijms150814475. Int J Mol Sci. 2014. PMID: 25141103 Free PMC article. Review.
-
Preclinical Evaluation of JAB-2485, a Potent AURKA Inhibitor with High Selectivity and Favorable Pharmacokinetic Properties.ACS Omega. 2024 May 3;9(19):21416-21425. doi: 10.1021/acsomega.4c01752. eCollection 2024 May 14. ACS Omega. 2024. PMID: 38764682 Free PMC article.
-
c-myc and N-myc promote active stem cell metabolism and cycling as architects of the developing brain.Oncotarget. 2010 Jun;1(2):120-30. doi: 10.18632/oncotarget.116. Oncotarget. 2010. PMID: 20651942 Free PMC article.
-
Calmodulin protects Aurora B on the midbody to regulate the fidelity of cytokinesis.Cell Cycle. 2013 Feb 15;12(4):663-73. doi: 10.4161/cc.23586. Epub 2013 Jan 31. Cell Cycle. 2013. PMID: 23370391 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

