A large and accurate collection of peptidase cleavages in the MEROPS database
- PMID: 20157488
- PMCID: PMC2790309
- DOI: 10.1093/database/bap015
A large and accurate collection of peptidase cleavages in the MEROPS database
Abstract
Peptidases are enzymes that hydrolyse peptide bonds in proteins and peptides. Peptidases are important in pathological conditions such as Alzheimer's disease, tumour and parasite invasion, and for processing viral polyproteins. The MEROPS database is an Internet resource containing information on peptidases, their substrates and inhibitors. The database now includes details of cleavage positions in substrates, both physiological and non-physiological, natural and synthetic. There are 39 118 cleavages in the collection; including 34 606 from a total of 10 513 different proteins and 2677 cleavages in synthetic substrates. The number of cleavages designated as 'physiological' is 13 307. The data are derived from 6095 publications. At least one substrate cleavage is known for 45% of the 2415 different peptidases recognized in the MEROPS database. The website now has three new displays: two showing peptidase specificity as a logo and a frequency matrix, the third showing a dynamically generated alignment between each protein substrate and its most closely related homologues. Many of the proteins described in the literature as peptidase substrates have been studied only in vitro. On the assumption that a physiologically relevant cleavage site would be conserved between species, the conservation of every site in terms of peptidase preference has been examined and a number have been identified that are not conserved. There are a number of cogent reasons why a site might not be conserved. Each poorly conserved site has been examined and a reason postulated. Some sites are identified that are very poorly conserved where cleavage is more likely to be fortuitous than of physiological relevance. This data-set is freely available via the Internet and is a useful training set for algorithms to predict substrates for peptidases and cleavage positions within those substrates. The data may also be useful for the design of inhibitors and for engineering novel specificities into peptidases.Database URL:http://merops.sanger.ac.uk.
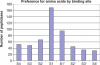
Figures


References
-
- Emanuelsson O, Brunak S, von Heijne G, et al. Locating proteins in the cell using TargetP, SignalP and related tools. Nat. Protoc. 2007;2:953–971. - PubMed
-
- Weissman AM. Regulating protein degradation by ubiquitination. Immunol. Today. 1997;18:189–198. - PubMed
-
- Drag M, Salvesen GS. DeSUMOylating enzymes—SENPs. IUBMB Life. 2008;60:734–742. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

