Dentinogenesis and Dentin Sialophosphoprotein (DSPP)
- PMID: 20157636
- PMCID: PMC2821091
- DOI: 10.2330/joralbiosci.51.134
Dentinogenesis and Dentin Sialophosphoprotein (DSPP)
Abstract
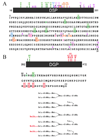
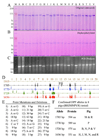
Dentin sialophosphoprotein (DSPP) is critical for proper mineralization of tooth dentin, and understanding its structure and function should yield important insights into how dentin biomineralization is controlled. During the recent six years, I have focused on characterizing DSPP-derived proteins isolated from developing porcine teeth. Porcine DSPP is expressed and secreted by odontoblasts and is processed by BMP-1, MMP-20 and MMP-2 into three main parts: dentin sialoprotein (DSP), dentin glycoprotein (DGP), and dentin phosphoprotein (DPP). We have learned that DSP is a proteoglycan that forms covalent dimers, DGP is a phosphorylated glycoprotein, and DPP is a highly phosphorylated intrinsically disordered protein that shows extensive length polymorphisms due to the genetic heterogeneity of its coding region.
Figures


Similar articles
-
Dentin Sialophophoprotein (DSPP) and Dentin.J Oral Biosci. 2008 Jan 1;50(1):33-44. doi: 10.2330/joralbiosci.50.33. J Oral Biosci. 2008. PMID: 20037676 Free PMC article.
-
Structural features, processing mechanism and gene splice variants of dentin sialophosphoprotein.Jpn Dent Sci Rev. 2018 Nov;54(4):183-196. doi: 10.1016/j.jdsr.2018.03.006. Epub 2018 Aug 25. Jpn Dent Sci Rev. 2018. PMID: 30302137 Free PMC article. Review.
-
Dentin sialophosphoprotein is processed by MMP-2 and MMP-20 in vitro and in vivo.J Biol Chem. 2006 Dec 15;281(50):38235-43. doi: 10.1074/jbc.M607767200. Epub 2006 Oct 17. J Biol Chem. 2006. PMID: 17046814
-
Dentin Sialophosphoprotein-derived Proteins in the Dental Pulp.J Dent Res. 2015 Aug;94(8):1120-7. doi: 10.1177/0022034515585715. Epub 2015 May 7. J Dent Res. 2015. PMID: 25951824
-
Dentin sialophosphoprotein-derived proteins in porcine pulp and dentin - Gene expression and function.J Oral Biosci. 2016 Nov;58(4):120-127. doi: 10.1016/j.job.2016.06.001. Epub 2016 Jul 7. J Oral Biosci. 2016. PMID: 32512680 Review.
Cited by
-
The role of thymosin beta 4 on odontogenic differentiation in human dental pulp cells.PLoS One. 2013 Apr 17;8(4):e61960. doi: 10.1371/journal.pone.0061960. Print 2013. PLoS One. 2013. Retraction in: PLoS One. 2025 Mar 3;20(3):e0320021. doi: 10.1371/journal.pone.0320021. PMID: 23613983 Free PMC article. Retracted.
-
The role of vitamin D receptor in predentin mineralization and dental repair after injury.Cell Tissue Res. 2024 Jun;396(3):343-351. doi: 10.1007/s00441-024-03886-7. Epub 2024 Mar 16. Cell Tissue Res. 2024. PMID: 38492000
-
Kappa-Carrageenan/Chitosan/Gelatin Scaffolds Provide a Biomimetic Microenvironment for Dentin-Pulp Regeneration.Int J Mol Sci. 2023 Mar 30;24(7):6465. doi: 10.3390/ijms24076465. Int J Mol Sci. 2023. PMID: 37047438 Free PMC article.
-
Hey1 functions as a positive regulator of odontogenic differentiation in odontoblast‑lineage cells.Int J Mol Med. 2018 Jan;41(1):331-339. doi: 10.3892/ijmm.2017.3254. Epub 2017 Nov 13. Int J Mol Med. 2018. PMID: 29138798 Free PMC article.
-
A pilot study on biological characteristics of human CD24(+) stem cells from the apical papilla.J Dent Sci. 2022 Jan;17(1):264-275. doi: 10.1016/j.jds.2021.01.012. Epub 2021 Apr 24. J Dent Sci. 2022. PMID: 35028047 Free PMC article.
References
-
- Linde A, Lussi A, Crenshaw MA. Mineral induction by immobilized polyanionic proteins. Calcif Tissue Int. 1989;44:286–295. - PubMed
-
- MacDougall M, Simmons D, Luan X, Nydegger J, Feng J, Gu TT. Dentin phosphoprotein and dentin sialoprotein are cleavage products expressed from a single transcript coded by a gene on human chromosome 4. Dentin phosphoprotein DNA sequence determination. J. Biol. Chem. 1997;272:835–842. - PubMed
-
- Qin C, Baba O, Butler WT. Post-translational modifications of sibling proteins and their roles in osteogenesis and dentinogenesis. Crit. Rev. Oral Biol. Med. 2004;15:126–136. - PubMed
-
- Veis A, Perry A. The phosphoprotein of the dentin matrix. Biochemistry. 1967;6:2409–2416. - PubMed
-
- Leaver AG, Triffitt JT, Holbrook IB. Newer knowledge of non-collagenous protein in dentin and cortical bone matrix. Clin Orthop Relat Res. 1975:269–292. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
