Promotion of exon 6 inclusion in HuD pre-mRNA by Hu protein family members
- PMID: 20159993
- PMCID: PMC2887941
- DOI: 10.1093/nar/gkq028
Promotion of exon 6 inclusion in HuD pre-mRNA by Hu protein family members
Abstract
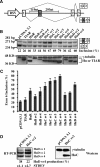
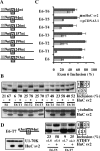
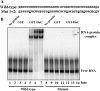
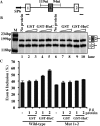
The Hu RNA-binding protein family consists of four members: HuR/A, HuB, HuC and HuD. HuR expression is widespread. The other three neuron-specific Hu proteins play an important role in neuronal differentiation through modulating multiple processes of RNA metabolism. In the splicing events examined previously, Hu proteins promote skipping of the alternative exons. Here, we report the first example where Hu proteins promote inclusion of an alternative exon, exon 6 of the HuD pre-mRNA. Sequence alignment analysis indicates the presence of several conserved AU-rich sequences both upstream and downstream to this alternatively spliced exon. We generated a human HuD exon 6 mini-gene reporter construct that includes these conserved sequences. Hu protein over-expression led to significantly increased exon 6 inclusion from this reporter and endogenous HuD. Studies using truncated and mutant HuD exon 6 reporters demonstrate that two AU-rich sequences located downstream of exon 6 are important. RNAi knockdown of Hu proteins decreased exon 6 inclusion. An in vitro splicing assay indicates that Hu proteins promote HuD exon 6 inclusion directly at the level of splicing. Our studies demonstrate that Hu proteins can function as splicing enhancers and expand the functional role of Hu proteins as splicing regulators.
Figures




Similar articles
-
Characterization of multiple exon 1 variants in mammalian HuD mRNA and neuron-specific transcriptional control via neurogenin 2.J Neurosci. 2012 Aug 15;32(33):11164-75. doi: 10.1523/JNEUROSCI.2247-12.2012. J Neurosci. 2012. PMID: 22895702 Free PMC article.
-
Regulation of neuron-specific alternative splicing of neurofibromatosis type 1 pre-mRNA.Mol Cell Biol. 2008 Feb;28(4):1240-51. doi: 10.1128/MCB.01509-07. Epub 2007 Dec 17. Mol Cell Biol. 2008. PMID: 18086893 Free PMC article.
-
ELAV/Hu RNA binding proteins determine multiple programs of neural alternative splicing.PLoS Genet. 2021 Apr 7;17(4):e1009439. doi: 10.1371/journal.pgen.1009439. eCollection 2021 Apr. PLoS Genet. 2021. PMID: 33826609 Free PMC article.
-
Role of HuD in nervous system function and pathology.Front Biosci (Schol Ed). 2013 Jan 1;5(2):554-63. doi: 10.2741/s389. Front Biosci (Schol Ed). 2013. PMID: 23277068 Review.
-
Diverse molecular functions of Hu proteins.Cell Mol Life Sci. 2008 Oct;65(20):3168-81. doi: 10.1007/s00018-008-8252-6. Cell Mol Life Sci. 2008. PMID: 18581050 Free PMC article. Review.
Cited by
-
ErbB4 alternative splicing mediates fetal mouse alveolar type II cell differentiation in vitro.Pediatr Res. 2022 Mar 25:10.1038/s41390-022-02013-y. doi: 10.1038/s41390-022-02013-y. Online ahead of print. Pediatr Res. 2022. PMID: 35338350 Free PMC article.
-
Dynamically expressed single ELAV/Hu orthologue elavl2 of bees is required for learning and memory.Commun Biol. 2021 Oct 28;4(1):1234. doi: 10.1038/s42003-021-02763-1. Commun Biol. 2021. PMID: 34711922 Free PMC article.
-
Neuronal Elav-like (Hu) proteins regulate RNA splicing and abundance to control glutamate levels and neuronal excitability.Neuron. 2012 Sep 20;75(6):1067-80. doi: 10.1016/j.neuron.2012.07.009. Neuron. 2012. PMID: 22998874 Free PMC article.
-
All three RNA recognition motifs and the hinge region of HuC play distinct roles in the regulation of alternative splicing.Nucleic Acids Res. 2013 May;41(9):5049-61. doi: 10.1093/nar/gkt166. Epub 2013 Mar 21. Nucleic Acids Res. 2013. PMID: 23525460 Free PMC article.
-
HuR function in disease.Front Biosci (Landmark Ed). 2012 Jan 1;17(1):189-205. doi: 10.2741/3921. Front Biosci (Landmark Ed). 2012. PMID: 22201738 Free PMC article. Review.
References
-
- Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 2008;40:1413–1415. - PubMed
-
- Black DL. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 2003;72:291–336. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

