Transcriptional corepressor TLE1 functions with Runx2 in epigenetic repression of ribosomal RNA genes
- PMID: 20160071
- PMCID: PMC2840145
- DOI: 10.1073/pnas.1000620107
Transcriptional corepressor TLE1 functions with Runx2 in epigenetic repression of ribosomal RNA genes
Abstract
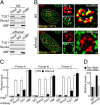
Epigenetic control of ribosomal RNA (rRNA) gene transcription by cell type-specific regulators, such as the osteogenic transcription factor Runx2, conveys cellular memory of growth and differentiation to progeny cells during mitosis. Here, we examined whether coregulatory proteins contribute to epigenetic functions that are mitotically transmitted by Runx2 in osteoblastic cells. We show that the transcriptional corepressor Transducin Like Enhancer-1 (TLE1) associates with rRNA genes during mitosis and interphase through interaction with Runx2. Mechanistically, depletion of TLE1 relieves Runx2-mediated repression of rRNA genes transcription and selectively increases histone modifications linked to active transcription. Biologically, loss of TLE-dependent rRNA gene repression coincides with increased global protein synthesis and enhanced cell proliferation. Our findings reinforce the epigenetic marking target genes by phenotypic transcription factors in mitosis and demonstrate a requirement for retention of coregulatory factors to sustain physiological control of gene expression during proliferation of lineage committed cells.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
A RUNX2-HDAC1 co-repressor complex regulates rRNA gene expression by modulating UBF acetylation.J Cell Sci. 2012 Jun 1;125(Pt 11):2732-9. doi: 10.1242/jcs.100909. Epub 2012 Mar 5. J Cell Sci. 2012. PMID: 22393235 Free PMC article.
-
Mitotic occupancy and lineage-specific transcriptional control of rRNA genes by Runx2.Nature. 2007 Jan 25;445(7126):442-6. doi: 10.1038/nature05473. Nature. 2007. PMID: 17251981
-
Subnuclear targeting of the Runx3 tumor suppressor and its epigenetic association with mitotic chromosomes.J Cell Physiol. 2009 Mar;218(3):473-9. doi: 10.1002/jcp.21630. J Cell Physiol. 2009. PMID: 19006109 Free PMC article.
-
Transcriptional co-repressors of Runx2.J Cell Biochem. 2006 May 1;98(1):54-64. doi: 10.1002/jcb.20805. J Cell Biochem. 2006. PMID: 16440320 Review.
-
Chromatin turn ons and turn offs of ribosomal RNA genes.Cell Cycle. 2004 Jul;3(7):880-3. Epub 2004 Jul 21. Cell Cycle. 2004. PMID: 15190210 Review.
Cited by
-
Deconstruction of the SS18-SSX fusion oncoprotein complex: insights into disease etiology and therapeutics.Cancer Cell. 2012 Mar 20;21(3):333-47. doi: 10.1016/j.ccr.2012.01.010. Cancer Cell. 2012. PMID: 22439931 Free PMC article.
-
Hyperphosphorylation by cyclin B/CDK1 in mitosis resets CUX1 DNA binding clock at each cell cycle.J Biol Chem. 2010 Oct 22;285(43):32834-32843. doi: 10.1074/jbc.M110.156406. Epub 2010 Aug 19. J Biol Chem. 2010. PMID: 20729212 Free PMC article.
-
Identification of GA-Binding Protein Transcription Factor Alpha Subunit (GABPA) as a Novel Bookmarking Factor.Int J Mol Sci. 2019 Mar 4;20(5):1093. doi: 10.3390/ijms20051093. Int J Mol Sci. 2019. PMID: 30836589 Free PMC article.
-
Inhibition of the RUNX1-CBFβ transcription factor complex compromises mammary epithelial cell identity: a phenotype potentially stabilized by mitotic gene bookmarking.Oncotarget. 2020 Jun 30;11(26):2512-2530. doi: 10.18632/oncotarget.27637. eCollection 2020 Jun 30. Oncotarget. 2020. PMID: 32655837 Free PMC article.
-
Reflections and perspectives on epigenetically mediated biological control: compromises in cancer and skeletal pathology.Acad Biol. 2025;3(2):10.20935/AcadBiol7628. doi: 10.20935/acadbiol7628. Epub 2025 Apr 11. Acad Biol. 2025. PMID: 40837125 Free PMC article.
References
-
- Enver T, Pera M, Peterson C, Andrews PW. Stem cell states, fates, and the rules of attraction. Cell Stem Cell. 2009;4:387–397. - PubMed
-
- Mellor J, Dudek P, Clynes D. A glimpse into the epigenetic landscape of gene regulation. Curr Opin Genet Dev. 2008;18:116–122. - PubMed
-
- Martínez-Balbás MA, Dey A, Rabindran SK, Ozato K, Wu C. Displacement of sequence-specific transcription factors from mitotic chromatin. Cell. 1995;83:29–38. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

