SRRM2, a potential blood biomarker revealing high alternative splicing in Parkinson's disease
- PMID: 20161708
- PMCID: PMC2817002
- DOI: 10.1371/journal.pone.0009104
SRRM2, a potential blood biomarker revealing high alternative splicing in Parkinson's disease
Abstract
Background: Parkinson's disease (PD) is a progressive neurodegenerative disorder that affects about five million people worldwide. Diagnosis remains clinical, based on phenotypic patterns. The discovery of laboratory markers that will enhance diagnostic accuracy, allow pre-clinical detection and tracking of disease progression is critically needed. These biomarkers may include transcripts with different isoforms.
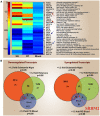
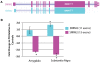
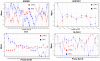
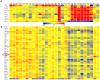
Methodology/principal findings: We performed extensive analysis on 3 PD microarray experiments available through GEO and found that the RNA splicing gene SRRM2 (or SRm300), sereine/arginine repetitive matrix 2, was the only gene differentially upregulated among all the three PD experiments. SRRM2 expression was not changed in the blood of other neurological diseased patients versus the healthy controls. Using real-time PCR, we report that the shorter transcript of SRRM2 was 1.7 fold (p = 0.008) upregulated in the substantia nigra of PDs vs controls while the longer transcript was 0.4 downregulated in both the substantia nigra (p = 0.03) and amygdala (p = 0.003). To validate our results and test for the possibility of alternative splicing in PD, we performed independent microarray scans, using Affymetrix Exon_ST1 arrays, from peripheral blood of 28 individuals (17 PDs and 11 Ctrls) and found a significant upregulation of the upstream (5') exons of SRRM2 and a downregulation of the downstream exons, causing a total of 0.7 fold down regulation (p = 0.04) of the long isoform. In addition, we report novel information about hundreds of genes with significant alternative splicing (differential exonic expression) in PD blood versus controls.
Conclusions/significance: The consistent dysregulation of the RNA splicing factor SRRM2 in two different PD neuronal sources and in PD blood but not in blood of other neurologically diseased patients makes SRRM2 a strong candidate gene for PD and draws attention to the role of RNA splicing in the disease.
Conflict of interest statement
Figures
References
-
- Papapetropoulos S, McCorquodale D. Gene Expression Profiling in Parkinson's disease: Discovery of valid biomarkers, molecular targets and biochemical pathways. Future Neurol. 2007;2:29–38.
-
- Grunblatt E, Mandel S, Jacob-Hirsch J, Zeligson S, Amariglo N, et al. Gene expression profiling of parkinsonian substantia nigra pars compacta; alterations in ubiquitin-proteasome, heat shock protein, iron and oxidative stress regulated proteins, cell adhesion/cellular matrix and vesicle trafficking genes. J Neural Transm. 2004;111:1543–1573. - PubMed
-
- Hauser MA, Li YJ, Xu H, Noureddine MA, Shao YS, et al. Expression profiling of substantia nigra in Parkinson disease, progressive supranuclear palsy, and frontotemporal dementia with parkinsonism. Arch Neurol. 2005;62:917–921. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

