Accelerated microevolution in an outer membrane protein (OMP) of the intracellular bacteria Wolbachia
- PMID: 20163713
- PMCID: PMC2843615
- DOI: 10.1186/1471-2148-10-48
Accelerated microevolution in an outer membrane protein (OMP) of the intracellular bacteria Wolbachia
Abstract
Background: Outer membrane proteins (OMPs) of Gram-negative bacteria are key players in the biology of bacterial-host interactions. However, while considerable attention has been given to OMPs of vertebrate pathogens, relatively little is known about the role of these proteins in bacteria that primarily infect invertebrates. One such OMP is found in the intracellular bacteria Wolbachia, which are widespread symbionts of arthropods and filarial nematodes. Recent experimental studies have shown that the Wolbachia surface protein (WSP) can trigger host immune responses and control cell death programming in humans, suggesting a key role of WSP for establishment and persistence of the symbiosis in arthropods.
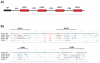
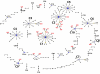
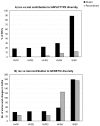
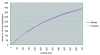
Results: Here we performed an analysis of 515 unique alleles found in 831 Wolbachia isolates, to investigate WSP structure, microevolution and population genetics. WSP shows an eight-strand transmembrane beta-barrel structure with four extracellular loops containing hypervariable regions (HVRs). A clustering approach based upon patterns of HVR haplotype diversity was used to group similar WSP sequences and to estimate the relative contribution of mutation and recombination during early stages of protein divergence. Results indicate that although point mutations generate most of the new protein haplotypes, recombination is a predominant force triggering diversity since the very first steps of protein evolution, causing at least 50% of the total amino acid variation observed in recently diverged proteins. Analysis of synonymous variants indicates that individual WSP protein types are subject to a very rapid turnover and that HVRs can accommodate a virtually unlimited repertoire of peptides. Overall distribution of WSP across hosts supports a non-random association of WSP with the host genus, although extensive horizontal transfer has occurred also in recent times.
Conclusions: In OMPs of vertebrate pathogens, large recombination impact, positive selection, reduced structural and compositional constraints, and extensive lateral gene transfer are considered hallmarks of evolution in response to the adaptive immune system. However, Wolbachia do not infect vertebrates. Here we predict that the rapid turnover of WSP loop motifs could aid in evading or inhibiting the invertebrate innate immune response. Overall, these features identify WSP as a strong candidate for future studies of host-Wolbachia interactions that affect establishment and persistence of this widespread endosymbiosis.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

