Alternative splicing of the histone demethylase LSD1/KDM1 contributes to the modulation of neurite morphogenesis in the mammalian nervous system
- PMID: 20164337
- PMCID: PMC6634524
- DOI: 10.1523/JNEUROSCI.5500-09.2010
Alternative splicing of the histone demethylase LSD1/KDM1 contributes to the modulation of neurite morphogenesis in the mammalian nervous system
Abstract
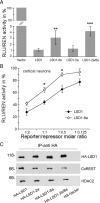
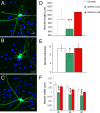
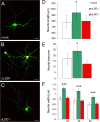
A variety of chromatin remodeling complexes are thought to orchestrate transcriptional programs that lead neuronal precursors from earliest commitment to terminal differentiation. Here we show that mammalian neurons have a specialized chromatin remodeling enzyme arising from a neurospecific splice variant of LSD1/KDM1, histone lysine specific demethylase 1, whose demethylase activity on Lys4 of histone H3 has been related to gene repression. We found that alternative splicing of LSD1 transcript generates four full-length isoforms from combinatorial retention of two identified exons: the 4 aa exon E8a is internal to the amine oxidase domain, and its inclusion is restricted to the nervous system. Remarkably, the expression of LSD1 splice variants is dynamically regulated throughout cortical development, particularly during perinatal stages, with a progressive increase of LSD1 neurospecific isoforms over the ubiquitous ones. Notably, the same LSD1 splice dynamics can be fairly recapitulated in cultured cortical neurons. Functionally, LSD1 isoforms display in vitro a comparable demethylase activity, yet the inclusion of the sole exon E8a reduces LSD1 repressor activity on a reporter gene. Additional distinction among isoforms is supported by the knockdown of neurospecific variants in cortical neurons resulting in the inhibition of neurite maturation, whereas overexpression of the same variants enhances it. Instead, perturbation of LSD1 isoforms that are devoid of the neurospecific exon elicits no morphogenic effect. Collectively, results demonstrate that the arousal of neuronal LSD1 isoforms pacemakes early neurite morphogenesis, conferring a neurospecific function to LSD1 epigenetic activity.
Figures





References
-
- Allis CD, Berger SL, Cote J, Dent S, Jenuwien T, Kouzarides T, Pillus L, Reinberg D, Shi Y, Shiekhattar R, Shilatifard A, Workman J, Zhang Y. New nomenclature for chromatin-modifying enzymes. Cell. 2007;131:633–636. - PubMed
-
- Ballas N, Battaglioli E, Atouf F, Andres ME, Chenoweth J, Anderson ME, Burger C, Moniwa M, Davie JR, Bowers WJ, Federoff HJ, Rose DW, Rosenfeld MG, Brehm P, Mandel G. Regulation of neuronal traits by a novel transcriptional complex. Neuron. 2001;31:353–365. - PubMed
-
- Ballas N, Grunseich C, Lu DD, Speh JC, Mandel G. REST and its corepressors mediate plasticity of neuronal gene chromatin throughout neurogenesis. Cell. 2005;121:645–657. - PubMed
-
- Barak O, Lazzaro MA, Cooch NS, Picketts DJ, Shiekhattar R. A tissue-specific, naturally occurring human SNF2L variant inactivates chromatin remodeling. J Biol Chem. 2004;279:45130–45138. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
