The major locus for mouse adenovirus susceptibility maps to genes of the hematopoietic cell surface-expressed LY6 family
- PMID: 20164425
- PMCID: PMC2832721
- DOI: 10.4049/jimmunol.0903363
The major locus for mouse adenovirus susceptibility maps to genes of the hematopoietic cell surface-expressed LY6 family
Abstract
Susceptibility to mouse adenovirus type 1 is associated with the major quantitative trait locus Msq1. Msq1 was originally mapped to a 13-Mb region of mouse chromosome (Chr) 15 in crosses between SJL/J and BALB/cJ inbred mice. We have now narrowed Msq1 to a 0.75-Mb interval from 74.68 to 75.43 Mb, defined by two anonymous markers, rs8259436 and D15Spn14, using data from 1396 backcross mice. The critical interval includes 14 Ly6 or Ly6-related genes, including Ly6a (encoding Sca-1/TAP), Ly6e (Sca-2/Tsa1), Ly6g (Gr-1), and gpihbp1 (GPI-anchored high-density lipoprotein-binding protein 1), as well as the gene encoding an aldosterone synthase (Cyp11b2). The Ly6 family members are attractive candidates for virus susceptibility genes because their products are GPI-anchored membrane proteins expressed on lymphoid and myeloid cells, with proposed functions in cell adhesion and cell signaling. To determine interstrain variation in susceptibility and produce additional resources for cloning Msq1, we assayed the susceptibility phenotype of four previously untested inbred mouse strains. Susceptibility of strain 129S6/SvEvTac was subsequently localized to the Ly6 complex region, using polymorphic genetic markers on Chr 15 in a population of 271 (129S6/SvEvTac x BALB/cJ)F(1) x BALB/cJ backcross mice. We identified a major 129S6/SvEvTac susceptibility allele, Msq1(129S6), on Chr 15 in the same region as Msq1(SJL). The results indicate that a major host factor in mouse adenovirus type 1 susceptibility is likely to be a member of the Ly6 gene family.
Figures
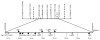
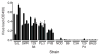

Similar articles
-
Contribution of a single host genetic locus to mouse adenovirus type 1 infection and encephalitis.mBio. 2012 May 29;3(3):e00131-12. doi: 10.1128/mBio.00131-12. Print 2012. mBio. 2012. PMID: 22647790 Free PMC article.
-
Polymorphisms in Ly6 genes in Msq1 encoding susceptibility to mouse adenovirus type 1.Mamm Genome. 2012 Apr;23(3-4):250-8. doi: 10.1007/s00335-011-9368-9. Epub 2011 Nov 20. Mamm Genome. 2012. PMID: 22101863 Free PMC article.
-
Identification of quantitative trait loci for susceptibility to mouse adenovirus type 1.J Virol. 2005 Sep;79(17):11517-22. doi: 10.1128/JVI.79.17.11517-11522.2005. J Virol. 2005. PMID: 16103204 Free PMC article.
-
Organization, evolution and functions of the human and mouse Ly6/uPAR family genes.Hum Genomics. 2016 Apr 21;10:10. doi: 10.1186/s40246-016-0074-2. Hum Genomics. 2016. PMID: 27098205 Free PMC article. Review.
-
Ly6 family proteins in neutrophil biology.J Leukoc Biol. 2013 Oct;94(4):585-94. doi: 10.1189/jlb.0113014. Epub 2013 Mar 29. J Leukoc Biol. 2013. PMID: 23543767 Review.
Cited by
-
Host genetic variation in susceptibility to Punta Toro virus.Virus Res. 2011 Apr;157(1):71-5. doi: 10.1016/j.virusres.2011.02.008. Epub 2011 Feb 12. Virus Res. 2011. PMID: 21320557 Free PMC article.
-
Contribution of a single host genetic locus to mouse adenovirus type 1 infection and encephalitis.mBio. 2012 May 29;3(3):e00131-12. doi: 10.1128/mBio.00131-12. Print 2012. mBio. 2012. PMID: 22647790 Free PMC article.
-
Matrix Metalloproteinase Activity in Infections by an Encephalitic Virus, Mouse Adenovirus Type 1.J Virol. 2017 Feb 28;91(6):e01412-16. doi: 10.1128/JVI.01412-16. Print 2017 Mar 15. J Virol. 2017. PMID: 28053109 Free PMC article.
-
Biochemistry and pathophysiology of intravascular and intracellular lipolysis.Genes Dev. 2013 Mar 1;27(5):459-84. doi: 10.1101/gad.209296.112. Genes Dev. 2013. PMID: 23475957 Free PMC article. Review.
-
Interferon-inducible LY6E Protein Promotes HIV-1 Infection.J Biol Chem. 2017 Mar 17;292(11):4674-4685. doi: 10.1074/jbc.M116.755819. Epub 2017 Jan 27. J Biol Chem. 2017. PMID: 28130445 Free PMC article.
References
-
- Brinton MA. Host susceptibility to viral disease. In: Nathanson N, Ahmed R, Gonzalez-Scarano R, Griffin DE, Holmes KV, Murphy FA, Robinson HL, editors. Viral Pathogenesis. Lippincott-Raven; Philadelphia: 1997. pp. 303–328.
-
- Mashimo T, Lucas M, Simon-Chazottes D, Frenkiel MP, Montagutelli X, Ceccaldi PE, Deubel V, Guénet JL, Desprès P. A nonsense mutation in the gene encoding 2′-5′-oligoadenylate synthetase/L1 isoform is associated with West Nile virus susceptibility in laboratory mice. Proc Natl Acad Sci USA. 2002;99:11311–11316. - PMC - PubMed
-
- Brown MG, Dokun AO, Heusel JW, Smith HRC, Beckman DL, Blattenberger EA, Dubbelde CE, Stone LR, Scalzo AA, Yokoyama WM. Vital involvement of a natural killer cell activation receptor in resistance to viral infection. Science. 2001;292:934–937. - PubMed
-
- Lee SH, Girard S, Macina D, Busa M, Zafer A, Belouchi A, Gros P, Vidal SM. Susceptibility to mouse cytomegalovirus is associated with deletion of an activating natural killer cell receptor of the C-type lectin superfamily. Nat Genet. 2001;28:42–45. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

