DomSVR: domain boundary prediction with support vector regression from sequence information alone
- PMID: 20165918
- PMCID: PMC2909371
- DOI: 10.1007/s00726-010-0506-6
DomSVR: domain boundary prediction with support vector regression from sequence information alone
Abstract
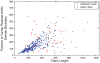
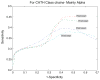
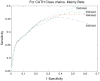
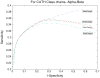
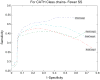
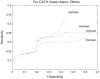
Protein domains are structural and fundamental functional units of proteins. The information of protein domain boundaries is helpful in understanding the evolution, structures and functions of proteins, and also plays an important role in protein classification. In this paper, we propose a support vector regression-based method to address the problem of protein domain boundary identification based on novel input profiles extracted from AAindex database. As a result, our method achieves an average sensitivity of approximately 36.5% and an average specificity of approximately 81% for multi-domain protein chains, which is overall better than the performance of published approaches to identify domain boundary. As our method used sequence information alone, our method is simpler and faster.
Figures




Similar articles
-
Improving protein structure similarity searches using domain boundaries based on conserved sequence information.BMC Struct Biol. 2009 May 19;9:33. doi: 10.1186/1472-6807-9-33. BMC Struct Biol. 2009. PMID: 19454035 Free PMC article.
-
Domain boundary prediction based on profile domain linker propensity index.Comput Biol Chem. 2006 Apr;30(2):127-33. doi: 10.1016/j.compbiolchem.2006.01.001. Epub 2006 Mar 13. Comput Biol Chem. 2006. PMID: 16531120
-
DoBo: Protein domain boundary prediction by integrating evolutionary signals and machine learning.BMC Bioinformatics. 2011 Feb 1;12:43. doi: 10.1186/1471-2105-12-43. BMC Bioinformatics. 2011. PMID: 21284866 Free PMC article.
-
DomHR: accurately identifying domain boundaries in proteins using a hinge region strategy.PLoS One. 2013 Apr 11;8(4):e60559. doi: 10.1371/journal.pone.0060559. Print 2013. PLoS One. 2013. PMID: 23593247 Free PMC article.
-
Improving the performance of DomainDiscovery of protein domain boundary assignment using inter-domain linker index.BMC Bioinformatics. 2006 Dec 18;7 Suppl 5(Suppl 5):S6. doi: 10.1186/1471-2105-7-S5-S6. BMC Bioinformatics. 2006. PMID: 17254311 Free PMC article.
Cited by
-
LigandRFs: random forest ensemble to identify ligand-binding residues from sequence information alone.BMC Bioinformatics. 2014;15 Suppl 15(Suppl 15):S4. doi: 10.1186/1471-2105-15-S15-S4. Epub 2014 Dec 3. BMC Bioinformatics. 2014. PMID: 25474163 Free PMC article.
-
Multi-head attention-based U-Nets for predicting protein domain boundaries using 1D sequence features and 2D distance maps.BMC Bioinformatics. 2022 Jul 19;23(1):283. doi: 10.1186/s12859-022-04829-1. BMC Bioinformatics. 2022. PMID: 35854211 Free PMC article.
-
TANGLE: two-level support vector regression approach for protein backbone torsion angle prediction from primary sequences.PLoS One. 2012;7(2):e30361. doi: 10.1371/journal.pone.0030361. Epub 2012 Feb 2. PLoS One. 2012. PMID: 22319565 Free PMC article.
-
DrugECs: An Ensemble System with Feature Subspaces for Accurate Drug-Target Interaction Prediction.Biomed Res Int. 2017;2017:6340316. doi: 10.1155/2017/6340316. Epub 2017 Jul 4. Biomed Res Int. 2017. PMID: 28744468 Free PMC article.
-
Protein domain identification methods and online resources.Comput Struct Biotechnol J. 2021 Feb 2;19:1145-1153. doi: 10.1016/j.csbj.2021.01.041. eCollection 2021. Comput Struct Biotechnol J. 2021. PMID: 33680357 Free PMC article. Review.
References
-
- Baldi P, Brunak S, Chauvin Y, Andersen CA, Nielsen H. Assessing the accuracy of prediction algorithms for classification: an overview. Bioinformatics. 2000;16:412–424. - PubMed
-
- Chen P, Wang B, Wong HS, Huang DS. Prediction of protein B-factors using multi-class bounded SVM. Protein Pept Lett. 2007;14(2):185–190. - PubMed
-
- Cheng J, Sweredoski MJ, Baldi P. DOMpro: protein domain prediction using profiles, secondary structure, relative solvent accessibility, and recursive neural networks. Data Min Knowl Discov. 2006;13:1–10.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

