Structural basis of membrane targeting by the Dock180 family of Rho family guanine exchange factors (Rho-GEFs)
- PMID: 20167601
- PMCID: PMC2857062
- DOI: 10.1074/jbc.M110.102517
Structural basis of membrane targeting by the Dock180 family of Rho family guanine exchange factors (Rho-GEFs)
Abstract
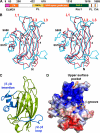
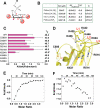
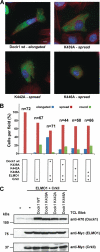
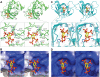
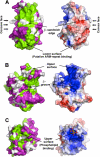
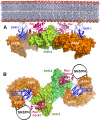
The Dock180 family of atypical Rho family guanine nucleotide exchange factors (Rho-GEFs) regulate a variety of processes involving cellular or subcellular polarization, including cell migration and phagocytosis. Each contains a Dock homology region-1 (DHR-1) domain that is required to localize its GEF activity to a specific membrane compartment where levels of phosphatidylinositol (3,4,5)-trisphosphate (PtdIns(3,4,5)P(3)) are up-regulated by the local activity of PtdIns 3-kinase. Here we define the structural and energetic bases of phosphoinositide specificity by the DHR-1 domain of Dock1 (a GEF for Rac1), and show that DHR-1 utilizes a C2 domain scaffold and surface loops to create a basic pocket on its upper surface for recognition of the PtdIns(3,4,5)P(3) head group. The pocket has many of the characteristics of those observed in pleckstrin homology domains. We show that point mutations in the pocket that abolish phospholipid binding in vitro ablate the ability of Dock1 to induce cell polarization, and propose a model that brings together recent mechanistic and structural studies to rationalize the central role of DHR-1 in dynamic membrane targeting of the Rho-GEF activity of Dock180.
Figures

Similar articles
-
GEF what? Dock180 and related proteins help Rac to polarize cells in new ways.Trends Cell Biol. 2007 Aug;17(8):383-93. doi: 10.1016/j.tcb.2007.05.001. Epub 2007 Aug 31. Trends Cell Biol. 2007. PMID: 17765544 Free PMC article. Review.
-
A minimal Rac activation domain in the unconventional guanine nucleotide exchange factor Dock180.Biochemistry. 2011 Feb 15;50(6):1070-80. doi: 10.1021/bi100971y. Epub 2011 Jan 20. Biochemistry. 2011. PMID: 21033699 Free PMC article.
-
Dissociation of Rac1(GDP).RhoGDI complexes by the cooperative action of anionic liposomes containing phosphatidylinositol 3,4,5-trisphosphate, Rac guanine nucleotide exchange factor, and GTP.J Biol Chem. 2008 Aug 8;283(32):22257-71. doi: 10.1074/jbc.M800734200. Epub 2008 May 27. J Biol Chem. 2008. PMID: 18505730 Free PMC article.
-
A novel and evolutionarily conserved PtdIns(3,4,5)P3-binding domain is necessary for DOCK180 signalling.Nat Cell Biol. 2005 Aug;7(8):797-807. doi: 10.1038/ncb1280. Epub 2005 Jul 17. Nat Cell Biol. 2005. PMID: 16025104 Free PMC article.
-
CZH proteins: a new family of Rho-GEFs.J Cell Sci. 2005 Nov 1;118(Pt 21):4937-46. doi: 10.1242/jcs.02671. J Cell Sci. 2005. PMID: 16254241 Review.
Cited by
-
GPCRs that Rhoar the Guanine nucleotide exchange factors.Small GTPases. 2022 Jan;13(1):84-99. doi: 10.1080/21541248.2021.1896963. Epub 2021 Apr 14. Small GTPases. 2022. PMID: 33849392 Free PMC article. Review.
-
Activation of Rac1 by Src-dependent phosphorylation of Dock180(Y1811) mediates PDGFRα-stimulated glioma tumorigenesis in mice and humans.J Clin Invest. 2011 Dec;121(12):4670-84. doi: 10.1172/JCI58559. Epub 2011 Nov 14. J Clin Invest. 2011. PMID: 22080864 Free PMC article.
-
Critical role of lipid membranes in polarization and migration of cells: a biophysical view.Biophys Rev. 2021 Jan 11;13(1):123-138. doi: 10.1007/s12551-021-00781-1. eCollection 2021 Feb. Biophys Rev. 2021. PMID: 33747247 Free PMC article. Review.
-
Deficiencies of the lipid-signaling enzymes phospholipase D1 and D2 alter cytoskeletal organization, macrophage phagocytosis, and cytokine-stimulated neutrophil recruitment.PLoS One. 2013;8(1):e55325. doi: 10.1371/journal.pone.0055325. Epub 2013 Jan 28. PLoS One. 2013. PMID: 23383154 Free PMC article.
-
Dendritic cells in inborn errors of immunity.Front Immunol. 2023 Jan 23;14:1080129. doi: 10.3389/fimmu.2023.1080129. eCollection 2023. Front Immunol. 2023. PMID: 36756122 Free PMC article. Review.
References
-
- Jaffe A. B., Hall A. (2005) Annu. Rev. Cell Dev. Biol. 21, 247–269 - PubMed
-
- Rossman K. L., Der C. J., Sondek J. (2005) Nat. Rev. Mol. Cell Biol. 6, 167–180 - PubMed
-
- Raftopoulou M., Hall A. (2004) Dev. Biol. 265, 23–32 - PubMed
-
- Côté J. F., Vuori K. (2002) J. Cell Sci. 115, 4901–4913 - PubMed
-
- Meller N., Irani-Tehrani M., Kiosses W. B., Del Pozo M. A., Schwartz M. A. (2002) Nat. Cell Biol. 4, 639–647 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

