Interleukin-7 compartmentalizes its receptor signaling complex to initiate CD4 T lymphocyte response
- PMID: 20167604
- PMCID: PMC2865298
- DOI: 10.1074/jbc.M110.104232
Interleukin-7 compartmentalizes its receptor signaling complex to initiate CD4 T lymphocyte response
Retraction in
-
Withdrawal: Interleukin-7 compartmentalizes its receptor signaling complex to initiate CD4 T lymphocyte response.J Biol Chem. 2019 Jan 25;294(4):1436. doi: 10.1074/jbc.W118.007350. J Biol Chem. 2019. PMID: 30808007 Free PMC article. No abstract available.
Abstract
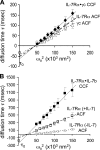
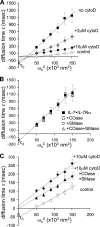
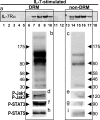
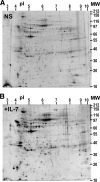
Interleukin (IL)-7 is a central cytokine that controls homeostasis of the CD4 T lymphocyte pool. Here we show on human primary cells that IL-7 binds to preassembled receptors made up of proprietary chain IL-7Ralpha and the common chain gammac shared with IL-2, -4, -9, -15, and -21 receptors. Upon IL-7 binding, both chains are driven in cholesterol- and sphingomyelin-rich rafts where associated signaling proteins Jak1, Jak3, STAT1, -3, and -5 are found to be phosphorylated. Meanwhile the IL-7.IL-7R complex interacts with the cytoskeleton that halts its diffusion as measured by single molecule fluorescence autocorrelated spectroscopy monitored by microimaging. Comparative immunoprecipitations of IL-7Ralpha signaling complex from non-stimulated and IL-7-stimulated cells confirmed recruitment of proteins such as STATs, but many others were also identified by mass spectrometry from two-dimensional gels. Among recruited proteins, two-thirds are involved in cytoskeleton and raft formation. Thus, early events leading to IL-7 signal transduction involve its receptor compartmentalization into membrane nanodomains and cytoskeleton recruitment.
Figures



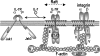
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

