DNA repair by the cryptic endonuclease activity of Mu transposase
- PMID: 20167799
- PMCID: PMC2890431
- DOI: 10.1073/pnas.0912615107
DNA repair by the cryptic endonuclease activity of Mu transposase
Abstract
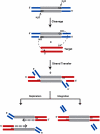
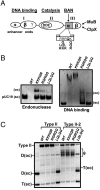
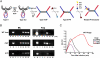
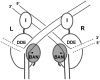
Phage Mu transposes by two distinct pathways depending on the specific stage of its life cycle. A common strand transfer intermediate is resolved differentially in the two pathways. During lytic growth, the intermediate is resolved by replication of Mu initiated within the flanking target DNA; during integration of infecting Mu, it is resolved without replication, by removal and repair of DNA from a previous host that is still attached to the ends of the incoming Mu genome. We have discovered that the cryptic endonuclease activity reported for the isolated C-terminal domain of the transposase MuA [Wu Z, Chaconas G (1995) A novel DNA binding and nuclease activity in domain III of Mu transposase: Evidence for a catalytic region involved in donor cleavage. EMBO J 14:3835-3843], which is not observed in the full-length protein or in the assembled transpososome in vitro, is required in vivo for removal of the attached host DNA or "5'flap" after the infecting Mu genome has integrated into the E. coli chromosome. Efficient flap removal also requires the host protein ClpX, which is known to interact with the C-terminus of MuA to remodel the transpososome for replication. We hypothesize that ClpX constitutes part of a highly regulated mechanism that unmasks the cryptic nuclease activity of MuA specifically in the repair pathway.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
ClpX and MuB interact with overlapping regions of Mu transposase: implications for control of the transposition pathway.Genes Dev. 1997 Jun 15;11(12):1561-72. doi: 10.1101/gad.11.12.1561. Genes Dev. 1997. PMID: 9203582
-
ClpX protein of Escherichia coli activates bacteriophage Mu transposase in the strand transfer complex for initiation of Mu DNA synthesis.EMBO J. 1996 Feb 15;15(4):935-44. EMBO J. 1996. PMID: 8631314 Free PMC article.
-
Mu transpososome and RecBCD nuclease collaborate in the repair of simple Mu insertions.Proc Natl Acad Sci U S A. 2014 Sep 30;111(39):14112-7. doi: 10.1073/pnas.1407562111. Epub 2014 Sep 2. Proc Natl Acad Sci U S A. 2014. PMID: 25197059 Free PMC article.
-
Remodeling protein complexes: insights from the AAA+ unfoldase ClpX and Mu transposase.Protein Sci. 2005 Aug;14(8):1945-54. doi: 10.1110/ps.051417505. Protein Sci. 2005. PMID: 16046622 Free PMC article. Review.
-
[Transposition as a way of existence: phage Mu].Genetika. 2003 May;39(5):637-56. Genetika. 2003. PMID: 12838611 Review. Russian.
Cited by
-
The μ transpososome structure sheds light on DDE recombinase evolution.Nature. 2012 Nov 15;491(7424):413-7. doi: 10.1038/nature11602. Epub 2012 Nov 7. Nature. 2012. PMID: 23135398 Free PMC article.
-
Mu insertions are repaired by the double-strand break repair pathway of Escherichia coli.PLoS Genet. 2012;8(4):e1002642. doi: 10.1371/journal.pgen.1002642. Epub 2012 Apr 12. PLoS Genet. 2012. PMID: 22511883 Free PMC article.
-
Repair of transposable phage Mu DNA insertions begins only when the E. coli replisome collides with the transpososome.Mol Microbiol. 2015 Aug;97(4):746-58. doi: 10.1111/mmi.13061. Epub 2015 Jun 6. Mol Microbiol. 2015. PMID: 25983038 Free PMC article.
-
A Well-Mixed E. coli Genome: Widespread Contacts Revealed by Tracking Mu Transposition.Cell. 2020 Feb 20;180(4):703-716.e18. doi: 10.1016/j.cell.2020.01.031. Epub 2020 Feb 13. Cell. 2020. PMID: 32059782 Free PMC article.
-
Universal platform for quantitative analysis of DNA transposition.Mob DNA. 2010 Nov 26;1(1):24. doi: 10.1186/1759-8753-1-24. Mob DNA. 2010. PMID: 21110848 Free PMC article.
References
-
- Symonds N, Toussaint A, Van de Putte P, Howe MM. Phage Mu. Plainview, NY: Cold Spring Harbor Laboratory Press; 1987.
-
- Akroyd JE, Symonds N. Evidence for a conservative pathway of transposition of bacteriophage Mu. Nature. 1983;303:84–86. - PubMed
-
- Harshey RM. Transposition without duplication of infecting bacteriophage Mu DNA. Nature. 1984;311:580–581. - PubMed
-
- Chaconas G, Harshey RM, Sarvetnick N, Bukhari AI. Predominant end products of prophage Mu DNA transposition during the lytic cycle are replicon fusions. J Mol Biol. 1981;150:341–359. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

