Mammalian recombination hot spots: properties, control and evolution
- PMID: 20168297
- PMCID: PMC4389181
- DOI: 10.1038/nrg2712
Mammalian recombination hot spots: properties, control and evolution
Abstract
Recombination, together with mutation, generates the raw material of evolution, is essential for reproduction and lies at the heart of all genetic analysis. Recent advances in our ability to construct genome-scale, high-resolution recombination maps and new molecular techniques for analysing recombination products have substantially furthered our understanding of this important biological phenomenon in humans and mice: from describing the properties of recombination hot spots in male and female meiosis to the recombination landscape along chromosomes. This progress has been accompanied by the identification of trans-acting systems that regulate the location and relative activity of individual hot spots.
Figures
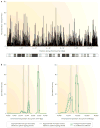
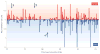
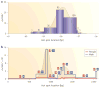
References
-
- Arnheim N, Calabrese P, Tiemann-Boege I. Mammalian meiotic recombination hot spots. Annu Rev Genet. 2007;41:369–399. - PubMed
-
- Buard J, de Massy B. Playing hide and seek with mammalian meiotic crossover hotspots. Trends Genet. 2007;23:301–309. - PubMed
-
- de Massy B. Distribution of meiotic recombination sites. Trends Genet. 2003;19:514–522. - PubMed
-
- Coop G, Przeworski M. An evolutionary view of human recombination. Nature Rev Genet. 2007;8:23–34. - PubMed
-
- Kauppi L, Jeffreys AJ, Keeney S. Where the crossovers are: recombination distributions in mammals. Nature Rev Genet. 2004;5:413–424. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

