Transcriptional targets of Drosophila JAK/STAT pathway signalling as effectors of haematopoietic tumour formation
- PMID: 20168330
- PMCID: PMC2838687
- DOI: 10.1038/embor.2010.1
Transcriptional targets of Drosophila JAK/STAT pathway signalling as effectors of haematopoietic tumour formation
Abstract
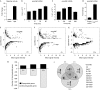
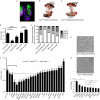
Although many signal transduction pathways have been implicated in the development of human disease, the identification of pathway targets and the biological processes that mediate disease progression remains challenging. One such disease-related pathway is the Janus kinase (JAK)/signal transducer and activator of transcription (STAT) cascade whose constitutive misactivation by the JAK2 V617F mutation underlies most human myeloproliferative disorders. Here, we use transcript profiling of Drosophila haemocyte-like cells to identify JAK/STAT target genes, combined with an in vivo model for JAK-induced blood cell overproliferation, to identify the main effectors required for haematopoietic tumour development. The identified human homologues of the Drosophila effectors were tested for potential V617F-mediated transcriptional regulation in human HeLa cells and compared with small interfering RNA-derived data, quantify their role in regulating the proliferation of cancer-derived cell lines. Such an inter-species approach is an effective way to identify factors with conserved functions that might be central to human disease.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

References
-
- Arbouzova NI, Zeidler MP (2006) JAK/STAT signalling in Drosophila: insights into conserved regulatory and cellular functions. Development 133: 2605–2616 - PubMed
-
- Dauer DJ, Ferraro B, Song L, Yu B, Mora L, Buettner R, Enkemann S, Jove R, Haura EB (2005) Stat3 regulates genes common to both wound healing and cancer. Oncogene 24: 3397–3408 - PubMed
-
- Dietzl G et al. (2007) A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila. Nature 448: 151–156 - PubMed
-
- Frey UH, Nuckel H, Sellmann L, Siemer D, Kuppers R, Durig J, Duhrsen U, Siffert W (2006) The GNAS1 T393C polymorphism is associated with disease progression and survival in chronic lymphocytic leukemia. Clin Cancer Res 12: 5686–5692 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

