Molecular poltergeists: mitochondrial DNA copies (numts) in sequenced nuclear genomes
- PMID: 20168995
- PMCID: PMC2820518
- DOI: 10.1371/journal.pgen.1000834
Molecular poltergeists: mitochondrial DNA copies (numts) in sequenced nuclear genomes
Abstract
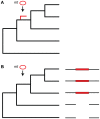
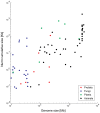
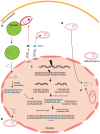
The natural transfer of DNA from mitochondria to the nucleus generates nuclear copies of mitochondrial DNA (numts) and is an ongoing evolutionary process, as genome sequences attest. In humans, five different numts cause genetic disease and a dozen human loci are polymorphic for the presence of numts, underscoring the rapid rate at which mitochondrial sequences reach the nucleus over evolutionary time. In the laboratory and in nature, numts enter the nuclear DNA via non-homolgous end joining (NHEJ) at double-strand breaks (DSBs). The frequency of numt insertions among 85 sequenced eukaryotic genomes reveal that numt content is strongly correlated with genome size, suggesting that the numt insertion rate might be limited by DSB frequency. Polymorphic numts in humans link maternally inherited mitochondrial genotypes to nuclear DNA haplotypes during the past, offering new opportunities to associate nuclear markers with mitochondrial markers back in time.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
References
-
- Gould SB, Waller RF, McFadden GI. Plastid evolution. Annu Rev Plant Biol. 2008;59:491–517. - PubMed
-
- Kleine T, Maier UG, Leister D. DNA transfer from organelles to the nucleus: the idiosyncratic genetics of endosymbiosis. Annu Rev Plant Biol. 2009;60:115–138. - PubMed
-
- Timmis JN, Ayliffe MA, Huang CY, Martin W. Endosymbiotic gene transfer: organelle genomes forge eukaryotic chromosomes. Nat Rev Genet. 2004;5:123–135. - PubMed
-
- Huang CY, Ayliffe MA, Timmis JN. Direct measurement of the transfer rate of chloroplast DNA into the nucleus. Nature. 2003;422:72–76. - PubMed
-
- Ricchetti M, Fairhead C, Dujon B. Mitochondrial DNA repairs double-strand breaks in yeast chromosomes. Nature. 1999;402:96–100. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

