Frequency and diversity of small cryptic plasmids in the genus Rahnella
- PMID: 20170524
- PMCID: PMC2831885
- DOI: 10.1186/1471-2180-10-56
Frequency and diversity of small cryptic plasmids in the genus Rahnella
Abstract
Background: Rahnella is a widely distributed genus belonging to the Enterobacteriaceae and frequently present on vegetables. Although Rahnella has interesting agro-economical and industrial properties and several strains possess antibiotic resistances and toxin genes which might spread within microbial communities, little is known about plasmids of this genus. Thus, we isolated a number of Rahnella strains and investigated their complements of small plasmids.
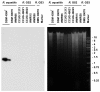
Results: In total 53 strains were investigated and 11 plasmids observed. Seven belonged to the ColE1 family; one was ColE2-like and three shared homology to rolling circle plasmids. One of them belonged to the pC194/pUB110 family and two showed similarity to poorly characterised plasmid groups. The G+C content of two rolling circle plasmids deviated considerably from that of Rahnella, indicating that their usual hosts might belong to other genera. Most ColE1-like plasmids formed a subgroup within the ColE1 family that seems to be fairly specific for Rahnella. Intriguingly, the multimer resolution sites of all ColE1-like plasmids had the same orientation with respect to the origin of replication. This arrangement might be necessary to prevent inappropriate synthesis of a small regulatory RNA that regulates cell division. Although the ColE1-like plasmids did not possess any mobilisation system, they shared large parts with high sequence identity in coding and non-coding regions. In addition, highly homologous regions of plasmids isolated from Rahnella and the chromosomes of Erwinia tasmaniensis and Photorhabdus luminescens could be identified.
Conclusions: For the genus Rahnella we observed plasmid-containing isolates at a frequency of 19%, which is in the average range for Enterobacteriaceae. These plasmids belonged to different groups with members of the ColE1-family most frequently found. Regions of striking sequence homology of plasmids and bacterial chromosomes highlight the importance of plasmids for lateral gene transfer (including chromosomal sequences) to distinct genera.
Figures





Similar articles
-
Isolation and characterization of pHW15, a small cryptic plasmid from Rahnella genomospecies 2.Plasmid. 2006 Nov;56(3):202-15. doi: 10.1016/j.plasmid.2006.05.007. Epub 2006 Jul 17. Plasmid. 2006. PMID: 16844220
-
Lateral transfer of rfb genes: a mobilizable ColE1-type plasmid carries the rfbO:54 (O:54 antigen biosynthesis) gene cluster from Salmonella enterica serovar Borreze.J Bacteriol. 1995 Sep;177(18):5247-53. doi: 10.1128/jb.177.18.5247-5253.1995. J Bacteriol. 1995. PMID: 7545154 Free PMC article.
-
Pseudomonas, the dominant polycyclic aromatic hydrocarbon-degrading bacteria isolated from Antarctic soils and the role of large plasmids in horizontal gene transfer.Environ Microbiol. 2006 Mar;8(3):455-65. doi: 10.1111/j.1462-2920.2005.00911.x. Environ Microbiol. 2006. PMID: 16478452
-
Rolling circle-replicating plasmids from gram-positive and gram-negative bacteria: a wall falls.Mol Microbiol. 1993 May;8(5):789-96. doi: 10.1111/j.1365-2958.1993.tb01625.x. Mol Microbiol. 1993. PMID: 8355606 Review.
-
Processing of plasmid DNA with ColE1-like replication origin.Plasmid. 2004 May;51(3):149-61. doi: 10.1016/j.plasmid.2003.12.002. Plasmid. 2004. PMID: 15109822 Review.
Cited by
-
PlasmidHunter: accurate and fast prediction of plasmid sequences using gene content profile and machine learning.Brief Bioinform. 2024 May 23;25(4):bbae322. doi: 10.1093/bib/bbae322. Brief Bioinform. 2024. PMID: 38960405 Free PMC article.
-
Emergence of plasmid stability under non-selective conditions maintains antibiotic resistance.Nat Commun. 2019 Jun 13;10(1):2595. doi: 10.1038/s41467-019-10600-7. Nat Commun. 2019. PMID: 31197163 Free PMC article.
-
Characterization of a Cryptic and Intriguing Low Molecular Weight Plasmid.Curr Microbiol. 2016 Mar;72(3):351-6. doi: 10.1007/s00284-015-0959-7. Epub 2015 Dec 15. Curr Microbiol. 2016. PMID: 26670037
-
Transcriptional responses to sucrose mimic the plant-associated life style of the plant growth promoting endophyte Enterobacter sp. 638.PLoS One. 2015 Jan 21;10(1):e0115455. doi: 10.1371/journal.pone.0115455. eCollection 2015. PLoS One. 2015. PMID: 25607953 Free PMC article.
-
Comparative Genomics Assisted Functional Characterization of Rahnella aceris ZF458 as a Novel Plant Growth Promoting Rhizobacterium.Front Microbiol. 2022 Apr 4;13:850084. doi: 10.3389/fmicb.2022.850084. eCollection 2022. Front Microbiol. 2022. PMID: 35444623 Free PMC article.
References
-
- Berge O, Heulin T, Achouak W, Richard C, Bally R, Balandreau J. Rahnella aquatilis, a nitrogen-fixing enteric bacterium associated with the rhizosphere of wheat and maize. Can J Microbiol. 1991;37:195–203.
-
- Heulin T, Berge O, Mavingui P, Gouzou L, Hebbar KP, Balandreau J. Bacillus polymyxa and Rahnella aquatilis, the dominant N2-fixing bacteria associated with wheat rhizosphere in French soils. Eur J Soil Biol. 1994;30:35–42.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

