Glucocorticoid receptor knock down reveals a similar apoptotic threshold but differing gene regulation patterns in T-cell and pre-B-cell acute lymphoblastic leukemia
- PMID: 20170710
- PMCID: PMC2844487
- DOI: 10.1016/j.mce.2010.02.014
Glucocorticoid receptor knock down reveals a similar apoptotic threshold but differing gene regulation patterns in T-cell and pre-B-cell acute lymphoblastic leukemia
Abstract
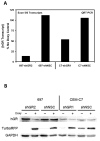
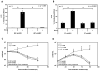
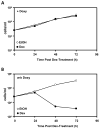
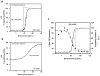
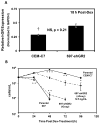
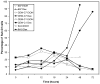
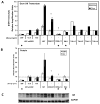
Glucocorticoids (GCs) are used in combination therapy for treating acute lymphoblastic leukemia (ALL). In T-cell (CEM-C7) and pre-B-cell (697) ALL cell lines, dexamethasone (Dex) treatment causes an auto-upregulation of glucocorticoid receptor (GR) mRNA transcripts and protein. We hypothesized that there is a threshold level of GR transcripts/protein needed for cells to respond to the apoptosis-inducing effects of hormone. GR knock down using a doxycycline-controllable shRNAmir indicated that the apoptotic response changes from sensitive to resistant with changing GR levels. Titration of the 697 cell GR to equal that of the CEM-C7 T-cell ALL line caused a shift in sensitivity to that seen in CEM-C7 cells. While the same level of GR is required to trigger apoptosis in both T-cell and pre-B-cell ALL lineages, similarities and differences were observed for the regulation of target genes in these lineages. These preliminary gene regulation patterns may lead to the development of a molecular signature for GC-sensitive and GC-resistant leukemia cells.
(c) 2010 Elsevier Ireland Ltd. All rights reserved.
Figures
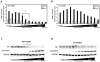

Similar articles
-
c-Myb interacts with the glucocorticoid receptor and regulates its level in pre-B-acute lymphoblastic leukemia cells.Mol Cell Endocrinol. 2012 Sep 25;361(1-2):124-32. doi: 10.1016/j.mce.2012.03.024. Epub 2012 Apr 10. Mol Cell Endocrinol. 2012. PMID: 22516378 Free PMC article.
-
A facile, branched DNA assay to quantitatively measure glucocorticoid receptor auto-regulation in T-cell acute lymphoblastic leukemia.Chin J Cancer. 2012 Aug;31(8):381-91. doi: 10.5732/cjc.012.10044. Epub 2012 Jun 26. Chin J Cancer. 2012. PMID: 22739263 Free PMC article.
-
A new, lineage specific, autoup-regulation mechanism for human glucocorticoid receptor gene expression in 697 pre-B-acute lymphoblastic leukemia cells.Mol Endocrinol. 2011 Jan;25(1):44-57. doi: 10.1210/me.2010-0249. Epub 2010 Nov 17. Mol Endocrinol. 2011. PMID: 21084380 Free PMC article.
-
Regulation of a distinctive set of genes in glucocorticoid-evoked apoptosis in CEM human lymphoid cells.Recent Prog Horm Res. 2003;58:175-97. doi: 10.1210/rp.58.1.175. Recent Prog Horm Res. 2003. PMID: 12795419 Review.
-
Resistance to glucocorticoid-induced apoptosis in lymphoblastic leukemia.J Endocrinol. 2003 Jul;178(1):19-27. doi: 10.1677/joe.0.1780019. J Endocrinol. 2003. PMID: 12844332 Review.
Cited by
-
c-Myb interacts with the glucocorticoid receptor and regulates its level in pre-B-acute lymphoblastic leukemia cells.Mol Cell Endocrinol. 2012 Sep 25;361(1-2):124-32. doi: 10.1016/j.mce.2012.03.024. Epub 2012 Apr 10. Mol Cell Endocrinol. 2012. PMID: 22516378 Free PMC article.
-
MicroRNAs and Glucocorticoid-Induced Apoptosis in Lymphoid Malignancies.ISRN Hematol. 2013;2013:348212. doi: 10.1155/2013/348212. Epub 2013 Jan 29. ISRN Hematol. 2013. PMID: 23431463 Free PMC article.
-
Research resource: transcriptional response to glucocorticoids in childhood acute lymphoblastic leukemia.Mol Endocrinol. 2012 Jan;26(1):178-93. doi: 10.1210/me.2011-1213. Epub 2011 Nov 10. Mol Endocrinol. 2012. PMID: 22074950 Free PMC article.
-
Thymocyte responsiveness to endogenous glucocorticoids is required for immunological fitness.J Clin Invest. 2012 Jul;122(7):2384-94. doi: 10.1172/JCI63067. Epub 2012 Jun 1. J Clin Invest. 2012. PMID: 22653054 Free PMC article.
-
A facile, branched DNA assay to quantitatively measure glucocorticoid receptor auto-regulation in T-cell acute lymphoblastic leukemia.Chin J Cancer. 2012 Aug;31(8):381-91. doi: 10.5732/cjc.012.10044. Epub 2012 Jun 26. Chin J Cancer. 2012. PMID: 22739263 Free PMC article.
References
-
- Abrams MT, Robertson NM, Yoon K, Wickstrom E. Inhibition of glucocorticoid-induced apoptosis by targeting the major splice variants of BIM mRNA with small interfering RNA and short hairpin RNA. J Biol Chem. 2004;279:55809–55817. - PubMed
-
- Ashraf J, Kunapuli S, Chilton D, Thompson EB. Cortivazol mediated induction of glucocorticoid receptor messenger ribonucleic acid in wild-type and dexamethasone-resistant human leukemic (CEM) cells. J Steroid Biochem Mol Biol. 1991;38:561–568. - PubMed
-
- Ashraf J, Thompson EB. Identification of the activation-labile gene: a single point mutation in the human glucocorticoid receptor presents as two distinct receptor phenotypes. Mol Endocrinol. 1993;7:631–642. - PubMed
-
- Bachmann PS, Gorman R, Papa RA, Bardell JE, Ford J, Kees UR, Marshall GM, Lock RB. Divergent mechanisms of glucocorticoid resistance in experimental models of pediatric acute lymphoblastic leukemia. Cancer Res. 2007;67:4482–4490. - PubMed
-
- Barrett TJ, Vig E, Vedeckis WV. Coordinate regulation of glucocorticoid receptor and c-jun gene expression is cell-type specific and exhibits differential hormonal sensitivity for down- and up-regulation. Biochemistry. 1996;35:9746–9753. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

