Glycoprotein gene sequence variation in rhesus monkey rhadinovirus
- PMID: 20172576
- PMCID: PMC2852144
- DOI: 10.1016/j.virol.2010.01.030
Glycoprotein gene sequence variation in rhesus monkey rhadinovirus
Abstract
Gene sequences for seven glycoproteins from 20 independent isolates of rhesus monkey rhadinovirus (RRV) and of the corresponding seven glycoprotein genes from nine strains of the Kaposi's sarcoma-associated herpesvirus (KSHV) were obtained and analyzed. Phylogenetic analysis revealed two discrete groupings of RRV gH sequences, two discrete groupings of RRV gL sequences and two discrete groupings of RRV gB sequences. We called these phylogenetic groupings gH(a), gH(b), gL(a), gL(b), gB(a) and gB(b). gH(a) was always paired with gL(a) and gH(b) was always paired with gL(b) for any individual RRV isolate. Since gH and gL are known to be interacting partners, these results suggest the need of matching sequence types for function of these cooperating proteins. gB phylogenetic grouping was not associated with gH/gL phylogenetic grouping. Our results demonstrate two distinct, distantly-related phylogenetic groupings of gH and gL of RRV despite a remarkable degree of sequence conservation within each individual phylogenetic group.
Copyright 2010 Elsevier Inc. All rights reserved.
Figures
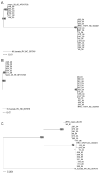




References
-
- Cawthon Lang KA. Primate Factsheets: Rhesus macaque (Macaca mulatta) Taxonomy, Mophology, & Ecology. 2005. Jul 20, http://pin.primate.wisc.edu/factsheets/entry/rhesus_macaque.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources

