Mass estimation of native proteins by blue native electrophoresis: principles and practical hints
- PMID: 20173216
- PMCID: PMC2953912
- DOI: 10.1074/mcp.M900526-MCP200
Mass estimation of native proteins by blue native electrophoresis: principles and practical hints
Abstract
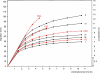
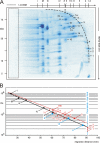
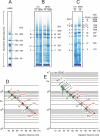
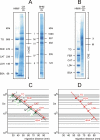
Blue native electrophoresis is one of the most popular techniques for mass estimation of native membrane proteins, but the use of non-optimal mass markers and acrylamide gels can compromise accuracy and reliability of the results. We present short protocols taking 10-30 min to prepare optimal sets of membrane protein markers from chicken, rat, mouse, and bovine heart. Especially heart materials from local supermarkets or butcher's shops, e.g. chicken or bovine heart, are ideal sources of high mass membrane protein standards. Considerable discrepancies between the migration behavior of membrane and soluble markers suggest using membrane protein markers for mass estimation of membrane proteins. Soluble standard proteins can be used, with some limitations, when soluble proteins are the focus. Principles and general rules for the determination of mass and oligomeric state of native membrane and soluble proteins are elaborated, and potential pitfalls are discussed.
Figures

Similar articles
-
Analysis of molecular masses and oligomeric states of protein complexes by blue native electrophoresis and isolation of membrane protein complexes by two-dimensional native electrophoresis.Anal Biochem. 1994 Mar;217(2):220-30. doi: 10.1006/abio.1994.1112. Anal Biochem. 1994. PMID: 8203750
-
LC-nanospray-MS/MS analysis of hydrophobic proteins from membrane protein complexes isolated by blue-native electrophoresis.J Mass Spectrom. 2005 Sep;40(9):1223-31. doi: 10.1002/jms.903. J Mass Spectrom. 2005. PMID: 16127664
-
A cost-effective method for simultaneous homo-oligomeric size determination and monodispersity conditions for membrane proteins.Anal Biochem. 2011 Sep 1;416(1):100-6. doi: 10.1016/j.ab.2011.05.007. Epub 2011 May 12. Anal Biochem. 2011. PMID: 21624344
-
Isolation of membrane protein complexes by blue native electrophoresis.Methods Mol Biol. 2008;424:423-31. doi: 10.1007/978-1-60327-064-9_33. Methods Mol Biol. 2008. PMID: 18369880 Review.
-
Features and applications of blue-native and clear-native electrophoresis.Proteomics. 2008 Oct;8(19):3974-90. doi: 10.1002/pmic.200800017. Proteomics. 2008. PMID: 18763698 Review.
Cited by
-
The small hydrophobic protein of the human respiratory syncytial virus forms pentameric ion channels.J Biol Chem. 2012 Jul 13;287(29):24671-89. doi: 10.1074/jbc.M111.332791. Epub 2012 May 23. J Biol Chem. 2012. PMID: 22621926 Free PMC article.
-
Native Gel Electrophoresis and Immunoblotting to Analyze Electron Transport Chain Complexes.Methods Mol Biol. 2021;2276:103-112. doi: 10.1007/978-1-0716-1266-8_7. Methods Mol Biol. 2021. PMID: 34060035
-
Penicillin-binding protein 5 can form a homo-oligomeric complex in the inner membrane of Escherichia coli.Protein Sci. 2011 Sep;20(9):1520-9. doi: 10.1002/pro.677. Epub 2011 Jul 13. Protein Sci. 2011. PMID: 21674665 Free PMC article.
-
Reconstitution of a minimal ESX-5 type VII secretion system suggests a role for PPE proteins in the outer membrane transport of proteins.mSphere. 2023 Oct 24;8(5):e0040223. doi: 10.1128/msphere.00402-23. Epub 2023 Sep 25. mSphere. 2023. PMID: 37747201 Free PMC article.
-
Ablation of mitochondrial DNA results in widespread remodeling of the mitochondrial complexome.EMBO J. 2021 Nov 2;40(21):e108648. doi: 10.15252/embj.2021108648. Epub 2021 Sep 20. EMBO J. 2021. PMID: 34542926 Free PMC article.
References
-
- Schägger H., von Jagow G. (1991) Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form. Anal. Biochem. 199, 223–231 - PubMed
-
- Schägger H., Cramer W. A., von Jagow G. (1994) Analysis of molecular masses and oligomeric states of protein complexes by blue native electrophoresis and isolation of membrane protein complexes by two-dimensional native electrophoresis. Anal. Biochem. 217, 220–230 - PubMed
-
- Wittig I., Braun H. P., Schägger H. (2006) Blue native PAGE. Nat. Protoc. 1, 418–428 - PubMed
-
- Wittig I., Schägger H. (2005) Advantages and limitations of clear native polyacrylamide gel electrophoresis. Proteomics 5, 4338–4346 - PubMed
-
- Wittig I., Karas M., Schägger H. (2007) High resolution clear-native electrophoresis for in-gel functional assays and fluorescence studies of membrane protein complexes. Mol. Cell. Proteomics 6, 1215–1225 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

