The X-ray crystal structure of Escherichia coli succinic semialdehyde dehydrogenase; structural insights into NADP+/enzyme interactions
- PMID: 20174634
- PMCID: PMC2823781
- DOI: 10.1371/journal.pone.0009280
The X-ray crystal structure of Escherichia coli succinic semialdehyde dehydrogenase; structural insights into NADP+/enzyme interactions
Abstract
Background: In mammals succinic semialdehyde dehydrogenase (SSADH) plays an essential role in the metabolism of the inhibitory neurotransmitter gamma-aminobutyric acid (GABA) to succinic acid (SA). Deficiency of SSADH in humans results in elevated levels of GABA and gamma-Hydroxybutyric acid (GHB), which leads to psychomotor retardation, muscular hypotonia, non-progressive ataxia and seizures. In Escherichia coli, two genetically distinct forms of SSADHs had been described that are essential for preventing accumulation of toxic levels of succinic semialdehyde (SSA) in cells.
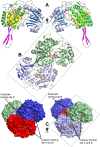
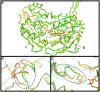
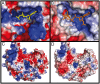
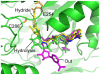
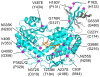
Methodology/principal findings: Here we structurally characterise SSADH encoded by the E coli gabD gene by X-ray crystallographic studies and compare these data with the structure of human SSADH. In the E. coli SSADH structure, electron density for the complete NADP+ cofactor in the binding sites is clearly evident; these data in particular revealing how the nicotinamide ring of the cofactor is positioned in each active site.
Conclusions/significance: Our structural data suggest that a deletion of three amino acids in E. coli SSADH permits this enzyme to use NADP+, whereas in contrast the human enzyme utilises NAD+. Furthermore, the structure of E. coli SSADH gives additional insight into human mutations that result in disease.
Conflict of interest statement
Figures



Similar articles
-
Succinic semialdehyde dehydrogenase: biochemical-molecular-clinical disease mechanisms, redox regulation, and functional significance.Antioxid Redox Signal. 2011 Aug 1;15(3):691-718. doi: 10.1089/ars.2010.3470. Epub 2011 Apr 10. Antioxid Redox Signal. 2011. PMID: 20973619 Free PMC article. Review.
-
Kinetic characterization and structural modeling of an NADP+-dependent succinic semialdehyde dehydrogenase from Anabaena sp. PCC7120.Int J Biol Macromol. 2018 Mar;108:615-624. doi: 10.1016/j.ijbiomac.2017.12.059. Epub 2017 Dec 11. Int J Biol Macromol. 2018. PMID: 29242124
-
Structural insight into the substrate inhibition mechanism of NADP(+)-dependent succinic semialdehyde dehydrogenase from Streptococcus pyogenes.Biochem Biophys Res Commun. 2015 Jun 5;461(3):487-93. doi: 10.1016/j.bbrc.2015.04.047. Epub 2015 Apr 16. Biochem Biophys Res Commun. 2015. PMID: 25888791
-
Saturation transfer difference NMR studies on substrates and inhibitors of succinic semialdehyde dehydrogenases.Biochem Biophys Res Commun. 2008 Aug 1;372(3):400-6. doi: 10.1016/j.bbrc.2008.04.183. Epub 2008 May 12. Biochem Biophys Res Commun. 2008. PMID: 18474219
-
Comparative genomics of aldehyde dehydrogenase 5a1 (succinate semialdehyde dehydrogenase) and accumulation of gamma-hydroxybutyrate associated with its deficiency.Hum Genomics. 2009 Jan;3(2):106-20. doi: 10.1186/1479-7364-3-2-106. Hum Genomics. 2009. PMID: 19164088 Free PMC article. Review.
Cited by
-
Succinic semialdehyde dehydrogenase: biochemical-molecular-clinical disease mechanisms, redox regulation, and functional significance.Antioxid Redox Signal. 2011 Aug 1;15(3):691-718. doi: 10.1089/ars.2010.3470. Epub 2011 Apr 10. Antioxid Redox Signal. 2011. PMID: 20973619 Free PMC article. Review.
-
Structure and function of phosphonoacetaldehyde dehydrogenase: the missing link in phosphonoacetate formation.Chem Biol. 2014 Jan 16;21(1):125-35. doi: 10.1016/j.chembiol.2013.11.006. Epub 2013 Dec 19. Chem Biol. 2014. PMID: 24361046 Free PMC article.
-
Crystallization and preliminary X-ray crystallographic studies of succinic semialdehyde dehydrogenase from Streptococcus pyogenes.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2012 Mar 1;68(Pt 3):288-91. doi: 10.1107/S1744309111052055. Epub 2012 Feb 22. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2012. PMID: 22442224 Free PMC article.
-
Gene replacement therapies for inherited disorders of neurotransmission: Current progress in succinic semialdehyde dehydrogenase deficiency.J Inherit Metab Dis. 2024 May;47(3):476-493. doi: 10.1002/jimd.12735. Epub 2024 Apr 6. J Inherit Metab Dis. 2024. PMID: 38581234 Free PMC article. Review.
-
Adaptive laboratory evolution accelerated glutarate production by Corynebacterium glutamicum.Microb Cell Fact. 2021 May 10;20(1):97. doi: 10.1186/s12934-021-01586-3. Microb Cell Fact. 2021. PMID: 33971881 Free PMC article.
References
-
- Chambliss K, Caudle DL, Hinson DD, Moomaw CR, Slaughter CA, et al. Molecular Cloning of the Mature NAD+-dependent Succinic Semialdehyde Dehydrogenase from Rat and Human. The Journal of Biological Chemistry. 1995;270:461–467. - PubMed
-
- Blaner WS, Churchich JE. The binding of NADH to succinic semialdehyde dehydrogenase. European journal of Biochemistry. 1980;109:431–437. - PubMed
-
- Chambliss K, Gibson KM. Succinic semialdehyde dehydrogenase from mammalian brain: subunit analysis using polyclonal antiserum. International journal of biochemistry & cell biology. 1992;24:1493–1499. - PubMed
-
- Lee B, Hong JW, Yoo BK, Lee SJ, Cho SW, et al. Bovine brain succinic semialdehyde dehydrogenase; purification, kinetics and reactivity of lysyl residues connected with catalytic activity. Molecules and cells. 1995;5:611–617.
-
- Ryzlak M, Pietruszko R. Human brain “high Km” aldehyde dehydrogenase: Purification, characterisation, and identification as NAD+ dependent succinic semialdehyde dehydrogenase. Archives of biochemistry and biophysics. 1988;266:386–396. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

