Restricted genetic diversity of HIV-1 subtype C envelope glycoprotein from perinatally infected Zambian infants
- PMID: 20174636
- PMCID: PMC2823783
- DOI: 10.1371/journal.pone.0009294
Restricted genetic diversity of HIV-1 subtype C envelope glycoprotein from perinatally infected Zambian infants
Erratum in
- PLoS One. 2010;5(3). doi: 10.1371/annotation/c2dd3548-93b7-480e-991a-93a728bca5fe
- PLoS One. 2010;5(3). doi: 10.1371/annotation/eadee58d-56c0-4399-b007-d80cc71e3f8a
Abstract
Background: Mother-to-child transmission of HIV-1 remains a significant problem in the resource-constrained settings where anti-retroviral therapy is still not widely available. Understanding the earliest events during HIV-1 transmission and characterizing the newly transmitted or founder virus is central to intervention efforts. In this study, we analyzed the viral env quasispecies of six mother-infant transmission pairs (MIPs) and characterized the genetic features of envelope glycoprotein that could influence HIV-1 subtype C perinatal transmission.
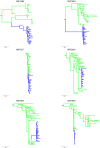
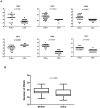
Methodology and findings: The V1-V5 region of env was amplified from 6 MIPs baseline samples and 334 DNA sequences in total were analyzed. A comparison of the viral population derived from the mother and infant revealed a severe genetic bottleneck occurring during perinatal transmission, which was characterized by low sequence diversity in the infant. Phylogenetic analysis indicates that most likely in all our infant subjects a single founder virus was responsible for establishing infection. Furthermore, the newly transmitted viruses from the infant had significantly fewer potential N-linked glycosylation sites in Env V1-V5 region and showed a propensity to encode shorter variable loops compared to the nontransmitted viruses. In addition, a similar intensity of selection was seen between mothers and infants with a higher rate of synonymous (dS) compared to nonsynonymous (dN) substitutions evident (dN/dS<1).
Conclusions: Our results indicate that a strong genetic bottleneck occurs during perinatal transmission of HIV-1 subtype C. This is evident through population diversity and phylogenetic patterns where a single viral variant appears to be responsible for infection in the infants. As a result the newly transmitted viruses are less diverse and harbored significantly less glycosylated envelope. This suggests that viruses with the restricted glycosylation in envelope glycoprotein appeared to be preferentially transmitted during HIV-1 subtype C perinatal transmission. In addition, our findings also indicated that purifying selection appears to predominate in shaping the early intrahost evolution of HIV-1 subtype C envelope sequences.
Conflict of interest statement
Figures


Similar articles
-
Comparative analysis of the fusion efficiency elicited by the envelope glycoprotein V1-V5 regions derived from human immunodeficiency virus type 1 transmitted perinatally.J Gen Virol. 2012 Dec;93(Pt 12):2635-2645. doi: 10.1099/vir.0.046771-0. Epub 2012 Sep 5. J Gen Virol. 2012. PMID: 22956734 Free PMC article.
-
Diversity of the human immunodeficiency virus type 1 (HIV-1) env sequence after vertical transmission in mother-child pairs infected with HIV-1 subtype A.J Virol. 2003 Mar;77(5):3050-7. doi: 10.1128/jvi.77.5.3050-3057.2003. J Virol. 2003. PMID: 12584330 Free PMC article.
-
The genetic bottleneck in vertical transmission of subtype C HIV-1 is not driven by selection of especially neutralization-resistant virus from the maternal viral population.J Virol. 2011 Aug;85(16):8253-62. doi: 10.1128/JVI.00197-11. Epub 2011 May 18. J Virol. 2011. PMID: 21593171 Free PMC article.
-
Mother-to-child transmission of HIV-1: advances and controversies of the twentieth centuries.AIDS Rev. 2004 Apr-Jun;6(2):67-78. AIDS Rev. 2004. PMID: 15332429 Review.
-
The role of HIV replicative fitness in perinatal transmission of HIV.Virol Sin. 2011 Jun;26(3):147-55. doi: 10.1007/s12250-011-3180-2. Epub 2011 Jun 12. Virol Sin. 2011. PMID: 21667335 Free PMC article. Review.
Cited by
-
HIV-1 maternal and infant variants show similar sensitivity to broadly neutralizing antibodies, but sensitivity varies by subtype.AIDS. 2013 Jun 19;27(10):1535-44. doi: 10.1097/QAD.0b013e32835faba5. AIDS. 2013. PMID: 23856624 Free PMC article.
-
Ultradeep single-molecule real-time sequencing of HIV envelope reveals complete compartmentalization of highly macrophage-tropic R5 proviral variants in brain and CXCR4-using variants in immune and peripheral tissues.J Neurovirol. 2018 Aug;24(4):439-453. doi: 10.1007/s13365-018-0633-5. Epub 2018 Apr 23. J Neurovirol. 2018. PMID: 29687407 Free PMC article.
-
No evidence of HIV replication in children on antiretroviral therapy.J Clin Invest. 2017 Oct 2;127(10):3827-3834. doi: 10.1172/JCI94582. Epub 2017 Sep 11. J Clin Invest. 2017. PMID: 28891813 Free PMC article. Clinical Trial.
-
Antibodies for prevention of mother-to-child transmission of HIV-1.Curr Opin HIV AIDS. 2015 May;10(3):177-82. doi: 10.1097/COH.0000000000000150. Curr Opin HIV AIDS. 2015. PMID: 25700205 Free PMC article. Review.
-
Viral Evolution and Cytotoxic T Cell Restricted Selection in Acute Infant HIV-1 Infection.Sci Rep. 2016 Jul 12;6:29536. doi: 10.1038/srep29536. Sci Rep. 2016. PMID: 27403940 Free PMC article.
References
-
- Luzuriaga K, Sullivan JL. Pediatric HIV-1 infection: advances and remaining challenges. AIDS Rev. 2002;4:21–26. - PubMed
-
- Scarlatti G, Leitner T, Halapi E, Wahlberg J, Marchisio P, et al. Comparison of variable region 3 sequences of human immunodeficiency virus type 1 from infected children with the RNA and DNA sequences of the virus populations of their mothers. Proc Natl Acad Sci U S A. 1993;90:1721–1725. - PMC - PubMed
-
- Wolinsky SM, Wike CM, Korber BT, Hutto C, Parks WP, et al. Selective transmission of human immunodeficiency virus type-1 variants from mothers to infants. Science. 1992;255:1134–1137. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

