Phylogenomics reveals a diverse Rickettsiales type IV secretion system
- PMID: 20176788
- PMCID: PMC2863512
- DOI: 10.1128/IAI.01384-09
Phylogenomics reveals a diverse Rickettsiales type IV secretion system
Abstract
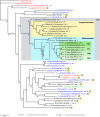
With an obligate intracellular lifestyle, Alphaproteobacteria of the order Rickettsiales have inextricably coevolved with their various eukaryotic hosts, resulting in small, reductive genomes and strict dependency on host resources. Unsurprisingly, large portions of Rickettsiales genomes encode proteins involved in transport and secretion. One particular transporter that has garnered recent attention from researchers is the type IV secretion system (T4SS). Homologous to the well-studied archetypal vir T4SS of Agrobacterium tumefaciens, the Rickettsiales vir homolog (rvh) T4SS is characterized primarily by duplication of several of its genes and scattered genomic distribution of all components in several conserved islets. Phylogeny estimation suggests a single event of ancestral acquirement of the rvh T4SS, likely from a nonalphaproteobacterial origin. Bioinformatics analysis of over 30 Rickettsiales genome sequences illustrates a conserved core rvh scaffold (lacking only a virB5 homolog), with lineage-specific diversification of several components (rvhB1, rvhB2, and rvhB9b), likely a result of modifications to cell envelope structure. This coevolution of the rvh T4SS and cell envelope morphology is probably driven by adaptations to various host cells, identifying the transporter as an important target for vaccine development. Despite the genetic intractability of Rickettsiales, recent advancements have been made in the characterization of several components of the rvh T4SS, as well as its putative regulators and substrates. While current data favor a role in effector translocation, functions in DNA uptake and release and/or conjugation cannot at present be ruled out, especially considering that a mechanism for plasmid transfer in Rickettsia spp. has yet to be proposed.
Figures



Similar articles
-
An anomalous type IV secretion system in Rickettsia is evolutionarily conserved.PLoS One. 2009;4(3):e4833. doi: 10.1371/journal.pone.0004833. Epub 2009 Mar 12. PLoS One. 2009. PMID: 19279686 Free PMC article.
-
Metagenome diversity illuminates the origins of pathogen effectors.mBio. 2024 May 8;15(5):e0075923. doi: 10.1128/mbio.00759-23. Epub 2024 Apr 2. mBio. 2024. PMID: 38564675 Free PMC article.
-
The Rickettsia type IV secretion system: unrealized complexity mired by gene family expansion.Pathog Dis. 2016 Aug;74(6):ftw058. doi: 10.1093/femspd/ftw058. Epub 2016 Jun 14. Pathog Dis. 2016. PMID: 27307105 Free PMC article.
-
Intracellular Rickettsiales: Insights into manipulators of eukaryotic cells.Trends Mol Med. 2011 Oct;17(10):573-83. doi: 10.1016/j.molmed.2011.05.009. Epub 2011 Jul 15. Trends Mol Med. 2011. PMID: 21763202 Review.
-
The Agrobacterium VirB/VirD4 T4SS: Mechanism and Architecture Defined Through In Vivo Mutagenesis and Chimeric Systems.Curr Top Microbiol Immunol. 2018;418:233-260. doi: 10.1007/82_2018_94. Curr Top Microbiol Immunol. 2018. PMID: 29808338 Free PMC article. Review.
Cited by
-
VirB10 vaccination for protection against Anaplasma phagocytophilum.BMC Microbiol. 2018 Dec 18;18(1):217. doi: 10.1186/s12866-018-1346-x. BMC Microbiol. 2018. PMID: 30563470 Free PMC article.
-
Recent molecular insights into rickettsial pathogenesis and immunity.Future Microbiol. 2013 Oct;8(10):1265-88. doi: 10.2217/fmb.13.102. Future Microbiol. 2013. PMID: 24059918 Free PMC article. Review.
-
Impact of Three Different Mutations in Ehrlichia chaffeensis in Altering the Global Gene Expression Patterns.Sci Rep. 2018 Apr 18;8(1):6162. doi: 10.1038/s41598-018-24471-3. Sci Rep. 2018. PMID: 29670161 Free PMC article.
-
Named entity recognition for bacterial Type IV secretion systems.PLoS One. 2011 Mar 29;6(3):e14780. doi: 10.1371/journal.pone.0014780. PLoS One. 2011. PMID: 21468321 Free PMC article.
-
Secretome of obligate intracellular Rickettsia.FEMS Microbiol Rev. 2015 Jan;39(1):47-80. doi: 10.1111/1574-6976.12084. Epub 2014 Dec 4. FEMS Microbiol Rev. 2015. PMID: 25168200 Free PMC article. Review.
References
-
- Aly, K. A., and C. Baron. 2007. The VirB5 protein localizes to the T-pilus tips in Agrobacterium tumefaciens. Microbiology 153:3766-3775. - PubMed
-
- Andersson, S. G., A. Zomorodipour, J. O. Andersson, T. Sicheritz-Ponten, U. C. Alsmark, R. M. Podowski, A. K. Naslund, A. S. Eriksson, H. H. Winkler, and C. G. Kurland. 1998. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396:133-140. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

