The endoplasmic reticulum Grp170 acts as a nucleotide exchange factor of Hsp70 via a mechanism similar to that of the cytosolic Hsp110
- PMID: 20177057
- PMCID: PMC2852982
- DOI: 10.1074/jbc.M109.096735
The endoplasmic reticulum Grp170 acts as a nucleotide exchange factor of Hsp70 via a mechanism similar to that of the cytosolic Hsp110
Abstract
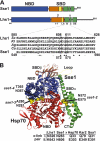
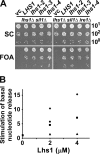
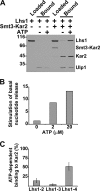
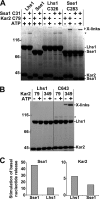
Grp170 and Hsp110 proteins constitute two evolutionary distinct branches of the Hsp70 family that share the ability to function as nucleotide exchange factors (NEFs) for canonical Hsp70s. Although the NEF mechanism of the cytoplasmic Hsp110s is well understood, little is known regarding the mechanism used by Grp170s in the endoplasmic reticulum. In this study, we compare the yeast Grp170 Lhs1 with the yeast Hsp110 Sse1. We find that residues important for Sse1 NEF activity are conserved in Lhs1 and that mutations in these residues in Lhs1 compromise NEF activity. As previously reported for Sse1, Lhs1 requires ATP to trigger nucleotide exchange in its cognate Hsp70 partner Kar2. Using site-specific cross-linking, we show that the nucleotide-binding domain (NBD) of Lhs1 interacts with the NBD of Kar2 face to face, and that Lhs1 contacts the side of the Kar2 NBD via its protruding C-terminal alpha-helical domain. To directly address the mechanism of nucleotide exchange, we have compared the hydrogen-exchange characteristics of a yeast Hsp70 NBD (Ssa1) in complex with either Sse1 or Lhs1. We find that Lhs1 and Sse1 induce very similar changes in the conformational dynamics in the Hsp70. Thus, our findings demonstrate that despite some differences between Hsp110 and Grp170 proteins, they use a similar mechanism to trigger nucleotide exchange.
Figures

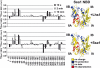
References
-
- Craig E. A. H. P. (2005) Protein Folding Handbook, pp. 490–515, Wiley, Weinheim, Germany
-
- Harrison C. J., Hayer-Hartl M., Di Liberto M., Hartl F., Kuriyan J. (1997) Science 276, 431–435 - PubMed
-
- Sondermann H., Scheufler C., Schneider C., Hohfeld J., Hartl F. U., Moarefi I. (2001) Science 291, 1553–1557 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

