Immunity and other defenses in pea aphids, Acyrthosiphon pisum
- PMID: 20178569
- PMCID: PMC2872881
- DOI: 10.1186/gb-2010-11-2-r21
Immunity and other defenses in pea aphids, Acyrthosiphon pisum
Abstract
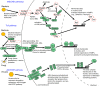
Background: Recent genomic analyses of arthropod defense mechanisms suggest conservation of key elements underlying responses to pathogens, parasites and stresses. At the center of pathogen-induced immune responses are signaling pathways triggered by the recognition of fungal, bacterial and viral signatures. These pathways result in the production of response molecules, such as antimicrobial peptides and lysozymes, which degrade or destroy invaders. Using the recently sequenced genome of the pea aphid (Acyrthosiphon pisum), we conducted the first extensive annotation of the immune and stress gene repertoire of a hemipterous insect, which is phylogenetically distantly related to previously characterized insects models.
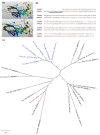
Results: Strikingly, pea aphids appear to be missing genes present in insect genomes characterized to date and thought critical for recognition, signaling and killing of microbes. In line with results of gene annotation, experimental analyses designed to characterize immune response through the isolation of RNA transcripts and proteins from immune-challenged pea aphids uncovered few immune-related products. Gene expression studies, however, indicated some expression of immune and stress-related genes.
Conclusions: The absence of genes suspected to be essential for the insect immune response suggests that the traditional view of insect immunity may not be as broadly applicable as once thought. The limitations of the aphid immune system may be representative of a broad range of insects, or may be aphid specific. We suggest that several aspects of the aphid life style, such as their association with microbial symbionts, could facilitate survival without strong immune protection.
Figures


Comment in
-
The pea aphid genome sequence brings theories of insect defense into question.Genome Biol. 2010;11(2):106. doi: 10.1186/gb-2010-11-2-106. Epub 2010 Feb 23. Genome Biol. 2010. PMID: 20236492 Free PMC article.
References
-
- Hufbauer RA. Aphid population dynamics: does resistance to parasitism influence population size? Ecol Entomol. 2002;27:25–32. doi: 10.1046/j.1365-2311.2002.0379a.x. - DOI

