The genetics of human adaptation: hard sweeps, soft sweeps, and polygenic adaptation
- PMID: 20178769
- PMCID: PMC2994553
- DOI: 10.1016/j.cub.2009.11.055
The genetics of human adaptation: hard sweeps, soft sweeps, and polygenic adaptation
Abstract
There has long been interest in understanding the genetic basis of human adaptation. To what extent are phenotypic differences among human populations driven by natural selection? With the recent arrival of large genome-wide data sets on human variation, there is now unprecedented opportunity for progress on this type of question. Several lines of evidence argue for an important role of positive selection in shaping human variation and differences among populations. These include studies of comparative morphology and physiology, as well as population genetic studies of candidate loci and genome-wide data. However, the data also suggest that it is unusual for strong selection to drive new mutations rapidly to fixation in particular populations (the 'hard sweep' model). We argue, instead, for alternatives to the hard sweep model: in particular, polygenic adaptation could allow rapid adaptation while not producing classical signatures of selective sweeps. We close by discussing some of the likely opportunities for progress in the field.
Copyright 2010 Elsevier Ltd. All rights reserved.
Figures
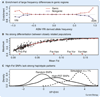
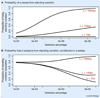

References
-
- Biswas S, Akey JM. Genomic insights into positive selection. Trends Genet. 2006;22:437–446. - PubMed
-
- Sabeti PC, Schaffner SF, Fry B, Lohmueller J, Varilly P, Shamovsky O, Palma A, Mikkelsen TS, Altshuler D, Lander ES. Positive natural selection in the human lineage. Science. 2006;312:1614–1620. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

