Reconstitution of eukaryotic lagging strand DNA replication
- PMID: 20178844
- PMCID: PMC2900510
- DOI: 10.1016/j.ymeth.2010.02.017
Reconstitution of eukaryotic lagging strand DNA replication
Abstract
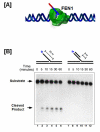
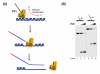
Eukaryotic DNA replication is a complex process requiring the proper functioning of a multitude of proteins to create error-free daughter DNA strands and maintain genome integrity. Even though synthesis and joining of Okazaki fragments on the lagging strand involves only half the DNA in the nucleus, the complexity associated with processing these fragments is about twice that needed for leading strand synthesis. Flap endonuclease 1 (FEN1) is the central component of the Okazaki fragment maturation pathway. FEN1 cleaves flaps that are displaced by DNA polymerase delta (pol delta), to create a nick that is effectively joined by DNA ligase I. The Pif1 helicase and Dna2 helicase/nuclease contribute to the maturation process by elongating the flap displaced by pol delta. Though the reason for generating long flaps is still a matter of debate, genetic studies have shown that Dna2 and Pif1 are both important components of DNA replication. Our current knowledge of the exact enzymatic steps that govern Okazaki fragment maturation has heavily derived from reconstitution reactions in vitro, which have augmented genetic information, to yield current mechanistic models. In this review, we describe both the design of specific DNA substrates that simulate intermediates of fragment maturation and protocols for reconstituting partial and complete lagging strand replication.
Copyright (c) 2010 Elsevier Inc. All rights reserved.
Figures





Similar articles
-
Dynamics of enzymatic interactions during short flap human Okazaki fragment processing by two forms of human DNA polymerase δ.DNA Repair (Amst). 2013 Nov;12(11):922-35. doi: 10.1016/j.dnarep.2013.08.008. Epub 2013 Sep 10. DNA Repair (Amst). 2013. PMID: 24035200 Free PMC article.
-
An alternative pathway for Okazaki fragment processing: resolution of fold-back flaps by Pif1 helicase.J Biol Chem. 2010 Dec 31;285(53):41712-23. doi: 10.1074/jbc.M110.146894. Epub 2010 Oct 19. J Biol Chem. 2010. PMID: 20959454 Free PMC article.
-
Flexibility of eukaryotic Okazaki fragment maturation through regulated strand displacement synthesis.J Biol Chem. 2008 Dec 5;283(49):34129-40. doi: 10.1074/jbc.M806668200. Epub 2008 Oct 16. J Biol Chem. 2008. PMID: 18927077 Free PMC article.
-
Polymerase dynamics at the eukaryotic DNA replication fork.J Biol Chem. 2009 Feb 13;284(7):4041-5. doi: 10.1074/jbc.R800062200. Epub 2008 Oct 3. J Biol Chem. 2009. PMID: 18835809 Free PMC article. Review.
-
Mechanism of Lagging-Strand DNA Replication in Eukaryotes.Adv Exp Med Biol. 2017;1042:117-133. doi: 10.1007/978-981-10-6955-0_6. Adv Exp Med Biol. 2017. PMID: 29357056 Review.
Cited by
-
DNA2-An Important Player in DNA Damage Response or Just Another DNA Maintenance Protein?Int J Mol Sci. 2017 Jul 18;18(7):1562. doi: 10.3390/ijms18071562. Int J Mol Sci. 2017. PMID: 28718810 Free PMC article. Review.
-
Targeting human papillomavirus genome replication for antiviral drug discovery.Antivir Ther. 2013;18(3):271-83. doi: 10.3851/IMP2612. Epub 2013 Apr 24. Antivir Ther. 2013. PMID: 23615820 Free PMC article. Review.
-
The flap endonuclease-1 promotes cellular tolerance to a chain-terminating nucleoside analog, alovudine, by counteracting the toxic effect of 53BP1.Nucleic Acids Res. 2025 Jul 8;53(13):gkaf617. doi: 10.1093/nar/gkaf617. Nucleic Acids Res. 2025. PMID: 40685548 Free PMC article.
-
Msh2-Msh3 interferes with Okazaki fragment processing to promote trinucleotide repeat expansions.Cell Rep. 2012 Aug 30;2(2):216-22. doi: 10.1016/j.celrep.2012.06.020. Epub 2012 Aug 2. Cell Rep. 2012. PMID: 22938864 Free PMC article.
-
DNA Polymerase Delta Exhibits Altered Catalytic Properties on Lysine Acetylation.Genes (Basel). 2023 Mar 23;14(4):774. doi: 10.3390/genes14040774. Genes (Basel). 2023. PMID: 37107532 Free PMC article.
References
-
- Waga S, Stillman B. The DNA replication fork in eukaryotic cells. Annu Rev Biochem. 1998;67:721–51. - PubMed
-
- Arezi B, Kuchta RD. Eukaryotic DNA primase. Trends Biochem Sci. 2000;25:572–6. - PubMed
-
- Tsurimoto T, Stillman B. Replication factors required for SV40 DNA replication in vitro. I. DNA structure-specific recognition of a primer-template junction by eukaryotic DNA polymerases and their accessory proteins. J Biol Chem. 1991;266:1950–60. - PubMed
-
- Burgers PM. Saccharomyces cerevisiae replication factor C. II. Formation and activity of complexes with the proliferating cell nuclear antigen and with DNA polymerases delta and epsilon. J Biol Chem. 1991;266:22698–706. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

